Using Machine Learning to Develop Personalized Vaccines for Cancer
A new tool combines multiple types of data to better support the development of vaccines that target patient-specific pathology.
Led by @virusesimmunity.bsky.social and @krishnaswamylab.bsky.social, a team of Yale researchers developed a new tool #Immunostruct that combines multiple types of data to better support the development of personalized vaccines, including vaccines for #cancer.
medicine.yale.edu/news-article...
27.02.2026 13:56
👍 0
🔁 1
💬 0
📌 0
We're @neuripsconf.bsky.social
HiPoNet: neurips.cc/virtual/2025...
HELM: Hyperbolic LLM openreview.net/forum?id=Rnb...
Talk: hyperboliclearning.github.io/events/neuri...
(Unireps/Neureps) Measure Before You Look openreview.net/forum?id=5Yn...
(FM4LS) CellSpliceNet: openreview.net/forum?id=gVA...
30.11.2025 18:44
👍 0
🔁 1
💬 0
📌 0
We have 3 papers at TAG-DS @NeurIPSConf!
DYMAG: Message Passing Using Dynamical-systems-based Waveforms (Oral) openreview.net/forum?id=WYi...
A Graph Laplacian Eigenvector Pretraining Method for GNNs (Spotlight) openreview.net/forum?id=r5c...
Reeb Graphs and Towers
openreview.net/forum?id=U8I...
30.11.2025 18:40
👍 1
🔁 1
💬 0
📌 0
(9/N) Grateful to lead authors Siddharth Viswanath and Hiren Madhu, and coauthors Dhananjay Bhaskar, Jake Kovalic, Dave Johnson, Chris Tape, Ian Adelstein, Rex Ying and Michael Perlmutter!
07.11.2025 14:19
👍 0
🔁 0
💬 1
📌 0
(8/N) ✨ In short, HiPoNet is a neural network that takes in a whole point cloud, and utilizes methods from geometry and topology to derive features of the point cloud.
→ Multiple learned views
→ Simplicial wavelets to extract multi-scale structure
07.11.2025 14:18
👍 0
🔁 0
💬 1
📌 0

(7/N) What’s more, HiPoNet’s learned feature weights are interpretable. For example, immune-related markers like CD11b, CD118, and FOXP3 consistently emerge as important across views, aligning with known tumor-immune biology.
07.11.2025 14:17
👍 0
🔁 0
💬 1
📌 0

(6/N) We find that HiPONet consistently outperforms other methods on these datasets.
07.11.2025 14:16
👍 0
🔁 0
💬 1
📌 0

(5/N) We then evaluate HiPOnet on patient level classification tasks using single cell and spatial proteomics data:
07.11.2025 14:15
👍 0
🔁 0
💬 1
📌 0

(4/N) We first check that HiPoNet preserves topological features of datasets by predicting persistence from the datasets:
07.11.2025 14:15
👍 0
🔁 0
💬 1
📌 0
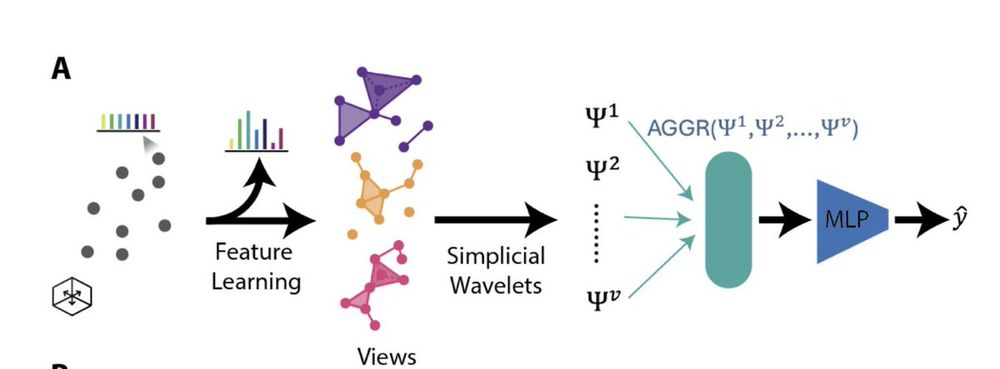
(3/N) HiPoNet models each cloud as a simplicial complex, and uses simplicial wavelet transforms to capture point-cloud level embeddings. Not only that---it uses multiview learning, i.e. different projections of the features to capture different underlying factors in the data.
07.11.2025 14:15
👍 0
🔁 0
💬 1
📌 0
(2/N) Modern technologies such as mass cytometry or scRNA-seq now allow large cohorts of patients to be measured. Each patient is actually a large single cell dataset creating a high-dimensional point cloud. But these point clouds are far more complex than 3D shapes that ML methods have handled..
07.11.2025 14:13
👍 0
🔁 0
💬 1
📌 0

(1/N) Thrilled to share that our paper HiPoNet (High dimensional Point cloud Network) to be presented at NeurIPS 2025! HiPoNet treats an entire high-dimensional point cloud as a datapoint! It captures multi-scale geometry and topology of the cloud perform classification and regression tasks.
07.11.2025 14:09
👍 12
🔁 3
💬 1
📌 0
Good morning from Vancouver! 🌞⛰️ We’re at #ICML2025 presenting at GenBio + MAS. Come chat with us about machine learning in biology!
With @RobertXiangruTang, @ChenLiu, @DanqiLiao. Views like this make the science even sweeter.🧬🧠
#ML #ComputationalBiology #ICML #Vancouver
17.07.2025 20:30
👍 1
🔁 1
💬 0
📌 0

Flyer for the Kavli Awards Research in Progress of June 23 showing pictures of the speakers in the Kavli logo, with a schedule:
12:00 pm Dhananjay Bhaskar, PhD
2024 Kavli Postdoctoral Fellowship
Krishnaswamy & Colón-Ramos labs
12:20 pm LaShae Nicholson, PhD
2022 Kavli Award for Academic Diversity
Strittmatter lab
12:40 pm José Jaime Martínez-Magaña, PhD
2023 Kavli Award for Academic Diversity
Montalvo-Ortiz lab
1:00 pm Kevin Chen, PhD
2024 Kavli Postdoctoral Fellowship
Emonet & Clark labs
1:20 pm break
1:30 pm Aaron Kuan, PhD &
Joerg Bewersdorf, PhD
2024 Kavli Teams Award
1:50 pm Enock Teefe, MD
2023 Kavli Award for Academic Diversity
Pittenger & Fernandez labs
2:10 pm Hongyan Hao, PhD
2024 Kavli Postdoctoral Fellowship
Hammarlund & De Camilli labs
Mark your calendars 🗓️ for June 23 for a Kavli Awards Research in Progress!🧠
Hear from Drs. Bhaskar (@krishnaswamylab.bsky.social), @lashaeneuroxp.bsky.social, Chen (@emonetlab.bsky.social), Martinez-Magaña (@janitzamontalvo.bsky.social), @kuanawanda.bsky.social, Teefe, & @haohy.bsky.social.
27.05.2025 20:31
👍 5
🔁 4
💬 0
📌 1

Accelerated learning of a noninvasive human brain-computer interface via manifold geometry
Brain-computer interfaces (BCIs) promise to restore and enhance a wide range of human capabilities. However, a barrier to the adoption of BCIs is how long it can take users to learn to control them. W...
New preprint! Excited to share our latest work “Accelerated learning of a noninvasive human brain-computer interface via manifold geometry” ft. outstanding former undergraduate Chandra Fincke, @glajoie.bsky.social, @krishnaswamylab.bsky.social, and @wutsaiyale.bsky.social's Nick Turk-Browne 1/8
03.04.2025 23:04
👍 66
🔁 20
💬 2
📌 3
Excited to organize this! Should be an exciting event!!
26.01.2025 13:55
👍 10
🔁 0
💬 0
📌 0
Great ideas to introduce to the community!! Enjoyed the talk!
25.01.2025 00:40
👍 4
🔁 0
💬 0
📌 0

Join us for an exciting session in @jointmath.bsky.social tomorrow! Here is our revised schedule!
10.01.2025 19:19
👍 15
🔁 1
💬 0
📌 0
We are at @jointmath.bsky.social Jan 8-11!
1/8: My talk at Topological Machine Learning I
1/8: Dhananjay Bhaskar at Contributed Paper session on Topology
1/11 join our special session on Mathematical and Computational Oncology
Also check out @pseudomanifold.topology.rocks on 1/9!!
08.01.2025 04:41
👍 9
🔁 3
💬 0
📌 0



Merry Xmas!
25.12.2024 15:15
👍 6
🔁 0
💬 0
📌 0

Join the Krishnaswamy Lab
We work on developing foundational mathematical <span class=
Looking for talented postdocs in deep learning for neuroscience and neuromodulation with the Murat Gunel lab in neurosurgery (medicine.yale.edu/lab/gunel/)! Projects include brain signal decoding, dynamics modeling for neuromodulation ASD and other conditions. See: krishnaswamylab.org/join
24.12.2024 15:05
👍 9
🔁 2
💬 0
📌 1
Frohe Weinachten!
24.12.2024 15:00
👍 1
🔁 0
💬 0
📌 0

(9/n) GSPA-Pt can be used to map patient sample manifolds. We mapped 48 melanoma patient scRNA-seq samples and classified response from the patient embedding using logistic regression. The GSPA-Pt gene embeddings achieved the highest classification performance.
21.12.2024 17:57
👍 3
🔁 0
💬 1
📌 0

(8/n) GSPA-LR concatenates ligand (L) and receptor (R) for a pair representation. LR pair modules captures a diverse range of LR profiles, finer the cluster-level analysis. For example, in skin cells, Module 5 includes Ccl5–Ccr5 link present in AG and AG CPI, epithelial, myeloid and T cells.
21.12.2024 17:55
👍 1
🔁 0
💬 1
📌 0

(7/n) With the Nik Joshi Lab at Yale we presented a new dataset of 39K CD8+ cells from LCMV infections. Interestingly only GSPA-based gene localization finds genes associated with type 1 interferon signaling. DE requires clustering, but DE genes in clusters do not reveal this signature!
21.12.2024 17:53
👍 1
🔁 0
💬 1
📌 0

(6/n) GSPA facilitates a measure called gene localization, via comparison to a uniform distribution. For PBMCs, cell embeddings using all genes versus only top localized genes showed similar structure, suggesting that localized genes capture cell-type variation and geometry.
21.12.2024 17:52
👍 1
🔁 0
💬 1
📌 0

(5/n) We show that for (PBMCs), GSPA with compressed wavelets grouped cell-type specific genes from PanglaoDB. In embryoid-body lineages early trajectory genes linked to embryonic stem cells (NANOG, POU5F1) and late trajectory genes linked to hemangioblast-specific signatures (CD34, PECAM1, TAL1).
21.12.2024 17:50
👍 1
🔁 0
💬 1
📌 0