At long last, my final PhD chapter is out: we developed a novel evolutionary simulator of bacterial pangenomes, Pansim, fitting it to data from >600K genomes using a likelihood-free framework, PopPUNK-mod, to explore neutral and adaptive pangenome dynamics www.biorxiv.org/content/10.6...
07.02.2026 10:08
👍 45
🔁 18
💬 2
📌 1
Merry X-SMASH!
My first first author paper is out on bioRxiv! 🖥
We present epsSMASH, a comprehensive and high-throughput tool for predicting exopolysaccharide (exoPS) biosynthetic gene clusters (BGCs) in bacterial genomes 🦠🧬🧫
1/8
24.12.2025 12:48
👍 57
🔁 21
💬 3
📌 3
Thrilled to share our new review online with
@scottvedwards.bsky.social in @Trends Ecol & Evo! “pangenomics is rapidly transforming our ability to dissect the genetic basis of ecological and evolutionary change in natural systems.”
26.12.2025 16:20
👍 13
🔁 6
💬 0
📌 0

Figure 1: (A) Anchor-based merging requires a common sequence (red) present in each partition. Multi-MUMs are merged by identifying overlaps between partition-specific matches in the anchor coordinate space, and a uniqueness threshold determines if a MUM is still unique in each partition after truncation. (B) String-based merging enables compu- tation of multi-MUMs between partitions without a common sequence. An example tree (left) is shown, highlighting the use case where partial multi-MUMs specific to internal nodes (starred) can be computed by merging subclade-based partitions up a tree. (right) MUM overlaps are computed by running Mumemto on the MUM sequences, and the uniqueness threshold array ensures overlaps remain unique across the merged dataset. (C) An example Burrows-Wheeler Transform (BWT), matrix (BWM), and Longest Com- mon Prefix (LCP) array, with sequence IDs for each suffix shown (ID). A non-maximal unique match (UM) is shown, and the uniqueness threshold for this match is found us- ing the flanking LCP values. (D) A partial multi-MUM (in blue) is found in all-but-one sequence (excluded in red). Using two anchor sequences (red and orange), all-but-one partial MUMs can be computed using an augmented anchor-based merging method (sec- tion 2.6).


Fantastic talk by @vikramshivakumar.bsky.social Mumemto—Scalable multi-MUM finding for pangenomes
Papers biorxiv.org/content/10.1101/2025.05.20.654611 & doi.org/10.1186/s13059-025-03644-0
Code: github.com/vikshiv/mume...
Very efficient pangenome visualization tool, revealing synteny and variations!
06.11.2025 01:13
👍 23
🔁 12
💬 1
📌 1

Labs in bacterial immunity
Hi everyone, a few years ago, we started a list of labs studying bacterial immunty for students, editors, conference organizers... (currently n=79).
Update time ! Send me a message to 1) add your lab or others 2) Correct info
docs.google.com/spreadsheets...
#Phagesky #Microsky
04.11.2025 15:06
👍 70
🔁 43
💬 13
📌 0

Non-conjugative plasmids limit their mobility to persist in nature
Sabnis et al. explain why non-conjugative plasmids move at a low rate in nature. While
increased mobility can easily evolve by incorporating phage DNA into plasmids, this
is disadvantageous because it...
Do plasmids really move around that much? Well, maybe not always
Thrilled to have contributed to this story with two of my favourite microbiologists: @jrpenades.bsky.social & @sanmillan.bsky.social
This great work was led by Akshay Sabnis & @wfigueroac3.bsky.social
www.cell.com/cell-reports...
22.10.2025 17:47
👍 38
🔁 17
💬 1
📌 0

Movi 2: Fast and Space-Efficient Queries on Pangenomes. #Pangenomes #SequenceQueries #Genomics #Bioinformatics @biorxiv-genomic.bsky.social 🧬 🖥️
www.biorxiv.org/content/10.1...
21.10.2025 13:49
👍 6
🔁 5
💬 0
📌 0

Text reads “Antibiotic resistance is rising,” with an upward arrow and a “40%” warning icon. Below, a text box says: “Between 2018 and 2023, resistance increased in over 40% of the monitored pathogen–antibiotic combinations, with an annual rise of 5–15%.”
Between 2018–2023, antibiotic resistance increased in over 40% of the pathogen–antibiotic combinations monitored, with an average annual rise of 5–15%.
Resistance is highest in the WHO South-East Asian & Eastern Mediterranean Regions, where 1 in 3 reported infections were resistant 👉 bit.ly/438Ta1u
15.10.2025 08:00
👍 102
🔁 52
💬 1
📌 7
#MicroSky
07.10.2025 12:58
👍 6
🔁 2
💬 0
📌 0
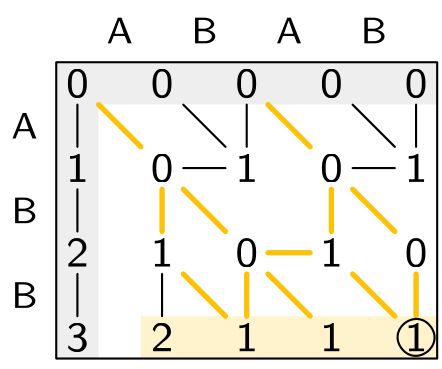
Sassy solves approximate string matching: finding all matches of a pattern in a text.
Sassy is out now!
Ever need to search for approximate matches of short DNA strings?
Sassy is the tool to use!
Available now wherever you get your code
With @rickbitloo.bsky.social
curiouscoding.nl/papers/sassy...
github.com/ragnarGrootK...
18.07.2025 20:20
👍 39
🔁 22
💬 2
📌 2

Phages with a broad host range are common across ecosystems - Nature Microbiology
Proximity-ligation-based sequencing from 111 samples and 5 environments reveals that a substantial proportion of phages infect multiple species.
In this great new paper, the authors use metagenomic Hi-C to show that #phage with broad host range are fairly common, contrary to the prevailing view on their biology. Cutting edge methods like Hi-C are shedding light on our blind spots in the microbial world.
29.09.2025 17:32
👍 11
🔁 3
💬 0
📌 0

Delighted to see our paper studying the evolution of plasmids over the last 100 years, now out! Years of work by Adrian Cazares, also Nick Thomson @sangerinstitute.bsky.social - this version much improved over the preprint. Final version should be open access, apols.
Thread 1/n
25.09.2025 21:28
👍 299
🔁 154
💬 14
📌 8

CompareM2 is a genomes-to-report pipeline for comparing microbial genomes. #MicrobialGenomes #Bioinformatics 🧬 🖥️
academic.oup.com/bioinformati...
18.09.2025 11:15
👍 5
🔁 3
💬 0
📌 0
Identification and Functional Insights into New Phage Tail-Like Bacteriocins (PTLBs) Targeting Pseudomonas aeruginosa as new antimicrobials https://www.biorxiv.org/content/10.1101/2025.08.20.671207v1
21.08.2025 04:16
👍 3
🔁 1
💬 0
📌 0

Pangenome analysis of transposable element insertion polymorphisms reveals features underlying cold tolerance in rice. #TransposableElements #TE #Pangenomes #RiceGenomes @natcomms.nature.com
www.nature.com/articles/s41...
17.08.2025 19:05
👍 23
🔁 7
💬 1
📌 0

Protein structure alignment significance is often exaggerated
Machine learning has generated millions of high-quality predicted protein structures, creating a need for computationally efficient structure search algorithms and robust estimates of statistical sign...
"We show that unrelated proteins have a universal tendency towards convergent evolution of secondary and tertiary motifs, causing an excess of high-scoring FP alignment... previous methods routinely overestimate significance by up to six orders of magnitude."
www.biorxiv.org/content/10.1...
17.08.2025 22:53
👍 52
🔁 23
💬 0
📌 2
The first paper of my PhD is now available as a preprint! 🎉
Transposable elements (TEs) don't just jump within fungal genomes, they also move extensively between species. In this study, we screened over 1,300 fungal genomes and found a conservative estimate of 5,500+ horizontal transfer events.
20.06.2025 19:39
👍 102
🔁 45
💬 7
📌 3






















