So happy to have been part of the @savitski-lab.bsky.social! I also want the sweater!
24.02.2026 16:34
👍 7
🔁 0
💬 0
📌 0

Our new preprint is out 🥳🥳🥳
Henipaviruses, like Nipah and Hendra, package their genomes inside helical shells built by thousands of nucleoproteins. These nucleocapsids are essential to protect the viral RNA, but how do they ever let the polymerase in to read the sequence?
👇
03.11.2025 12:26
👍 111
🔁 47
💬 3
📌 2

Mosquitoes like Aedes aegypti spread viruses that cause major global diseases. We decided to have a closer look at a family of mosquito carbohydrate-binding proteins involved in infecting these vector species. We found some unexpected things 😁
Preprint out now 🦟🦟🦟
www.biorxiv.org/content/10.6...
13.02.2026 09:04
👍 33
🔁 17
💬 1
📌 1
ICYMI during the holidays. I am really proud of Bolor's efforts and the first work coming out of the lab!
Read it here: www.biorxiv.org/content/10.6...
07.01.2026 08:13
👍 8
🔁 1
💬 0
📌 0
New paper from the lab!
Led by Patrick Müller: we show that non-antibiotic compounds — but not macrolide antibiotics — can induce an efflux pump that protects Bacteroidaceae from macrolides.
A neat twist on drug–microbe interactions... www.tandfonline.com/doi/full/10....
09.12.2025 10:33
👍 19
🔁 11
💬 0
📌 1

The pcnB gene sustains Shigella flexneri virulence
Author summary Bacterial infections represent a major global threat. Understanding the genetic determinants promoting infections is crucial to overcome this threat. Shigella is an intracellular bacter...
Thrilled to share that our study on how the pcnB gene sustains Shigella virulence is now published in PLOS Pathogens: journals.plos.org/plospathogen...
In short: pcnB ➝increased virulence-plasmid replication ➝ elevated T3SS expression ➝ enhanced invasion/spread within the intestinal epithelium.
21.11.2025 11:28
👍 14
🔁 7
💬 2
📌 0
#ProudPI
27.06.2025 13:06
👍 6
🔁 0
💬 0
📌 0
New preprint out: www.biorxiv.org/content/10.1...
We followed up on our recent Nature Genetics paper mapping Shigella colonization in intestinal organoids with a deep dive into one of the screen hits: pcnB.
#Shigella #virulence #microsky
17.06.2025 09:35
👍 4
🔁 3
💬 1
📌 0
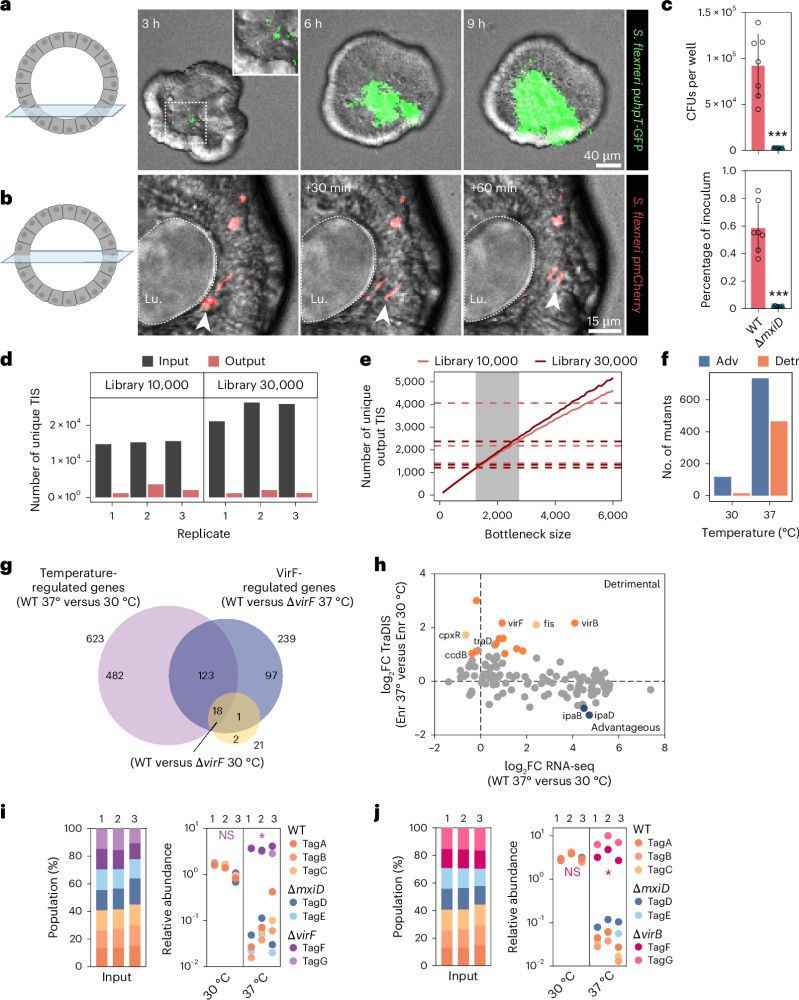
A scalable gut epithelial organoid model reveals the genome-wide colonization landscape of a human-adapted pathogen
Nature Genetics - A genome-wide screen using human gut epithelial organoids combined with transposon-directed insertion sequencing identifies over 100 Shigella flexneri genes required for...
Our latest work combines human gut epithelial #organoid culture with experimental #Shigella #infections, transposon mutagenesis, and statistical modeling. This has led to the mapping of the comprehensive geneset that drives Shigella epithelial colonization.
Out @natgenet.nature.com:
rdcu.be/eqGgf
12.06.2025 14:14
👍 32
🔁 20
💬 3
📌 1
Do we really know the antibiotics that we already have?
Check out our story on 60y old drug nitroxoline and its new tricks!
@goetheuni.bsky.social @embl.org
www.nature.com/articles/s41...
+ great piece from the synchrotron @esrf.fr where we chased nx effects on metals
www.esrf.fr/home/news/ge...
24.04.2025 07:16
👍 15
🔁 5
💬 3
📌 2
Looking forward to speaking in this webinar series next Wednesday.
16.04.2025 17:46
👍 4
🔁 1
💬 0
📌 0
So happy to see the first paper on research from the lab out! Just a review for now, but it gives an idea of the avenues we are exploring.
15.04.2025 11:57
👍 11
🔁 2
💬 0
📌 0
Looking forward to sharing with this community what our lab is doing
21.01.2025 08:30
👍 4
🔁 0
💬 1
📌 0








