The moment you’ve all been waiting for…
🦠 SAVE THE DATE! 🦠
BBM2026 will be held from June 22nd - 23rd at Boston University’s George Sherman Union. Our featured speaker this year is Dr. Eric Skaar from Vanderbilt University!
Registration opens soon!
More info at: bostonbacterial.org
#BBM2026
12.02.2026 11:37
👍 21
🔁 17
💬 1
📌 0

Model for Clostridioides difficile germination. Top: Model of the C. difficile proteins required for germination during sporulation. From left to right: interdomain processing of the CspBA fusion protein by the protease YabG occurs in the spore coat layer; heterodimerization of CspC and CspA and homodimerization of CspB; and loading of key germination proteins into the cortex region of the mature spore. Bottom right: Model of a C. difficile spore undergoing germination. Sensing of germinant and co-germinant signals (either directly or indirectly) by the CspC:CspA heterodimer (1) leads to the heterodimer adopting an active conformation (2). Bottom left: CspC:CspA heterodimer space-filling model. Residues at the center of the CspC:CspA heterodimer binding interface (I.) play important roles in transducing germinant and co-germinant signals, while residues at the periphery of the binding interface (II.) help stabilize the complex. Signal transduction by the CspC:CspA heterodimer initiates the activation of CspB (either directly or indirectly) (3). Active CspB then proteolytically activates the cortex-lytic enzyme SleC (4), which will go on to degrade the spore cortex (5), allowing for rehydration of the spore core and resumption of metabolic activity.
Unlike other #bacteria, C. difficile must detect both germinant & co-germinant signals to trigger #spore #germination. This study finds that the CspC:CspA complex is a key signaling node that integrates environmental cues to regulate #Cdifficile spore germination @plosbiology.org 🧪 plos.io/4rtCKdZ
03.02.2026 09:20
👍 6
🔁 3
💬 1
📌 2
Great new story from Sophie Helaine and Molly Sargen!
www.helainelab.com
28.01.2026 23:01
👍 35
🔁 18
💬 0
📌 0

The #SubtiWiki Paper of the month for June 2025 has been selected.
Congratulations, @apbrogan.bsky.social @ @harvardmed.bsky.social
@natmicrobiol.nature.com
subtiwiki.uni-goettingen.de/wiki/index.p...
30.06.2025 12:50
👍 6
🔁 3
💬 0
📌 0
Thank you so much!
19.06.2025 03:09
👍 0
🔁 0
💬 0
📌 0
Thanks Adrienne!
18.06.2025 00:58
👍 0
🔁 0
💬 0
📌 0
Thank you! and will do.
17.06.2025 14:34
👍 1
🔁 0
💬 0
📌 0
Thank you Jörg! I was so inspired by your work over the years.
17.06.2025 14:33
👍 1
🔁 0
💬 1
📌 0

Cyclic-di-AMP modulates cellular turgor
in response to defects in bacterial cell wall
synthesis

Cyclic-di-AMP modulates cellular turgor
in response to defects in bacterial cell wall
synthesis
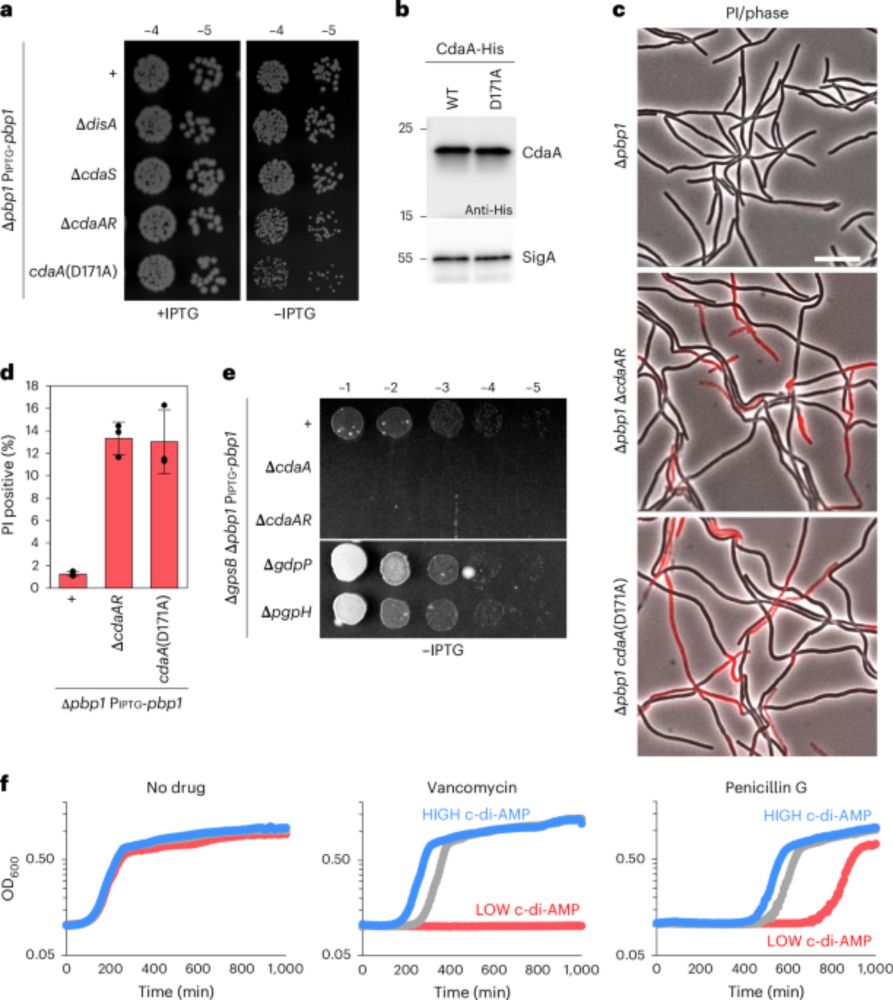
Brogan et al uncover a signaling pathway in which levels of the nucleotide second messenger c-di-AMP increase in response to defects in cell wall synthesis. This regulatory pathway decreases the cytoplasmic turgor pressure and protects the cell from lysis: www.nature.com/articles/s41...
17.06.2025 13:01
👍 14
🔁 4
💬 0
📌 0
This was a great collaboration with Dr. Paola Bardetti and Dr. Rico Rojas at NYU. With their help, we provide evidence that c-di-AMP functions to control the cytoplasmic turgor pressure. The c-di-AMP mediated reduction in turgor protects cells from lysis.
17.06.2025 12:20
👍 1
🔁 0
💬 0
📌 0
In short, there is: the c-di-AMP cyclase/regulator complex CdaAR upregulates cyclic-di-AMP in response to defects in cell wall synthesis. Similar to other envelope stress response systems in B. subtills, the cyclase regulator CdaR uses an intrinsically disordered region to sense cell wall defects.
17.06.2025 12:20
👍 2
🔁 0
💬 1
📌 0
c-di-AMP has long been implicated in resistance to cell wall targeting antibiotics. We hypothesized that there could be a regulatory connection between the cell envelope and c-di-AMP levels.
17.06.2025 12:20
👍 0
🔁 0
💬 1
📌 0
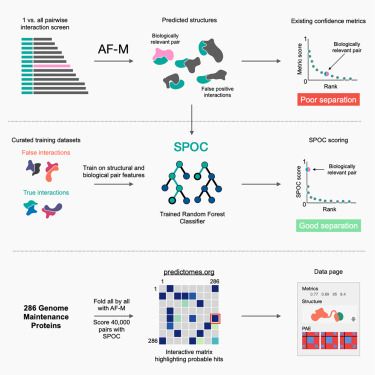
🚨👉 Please check our recent work on bacterial cell division. In situ Cryo-ET reveals the cellular function of the penicillin binding protein 1b supported by AFM, live-cell imaging, in silico AlphaFold proteome screen and TIRFM. Hope you enjoy the read! #teamtomo #cryo-ET ❄️🔬🐎 big thanks to the team!
03.04.2025 09:08
👍 56
🔁 23
💬 0
📌 4

SAVE THE DATE ‼️ 📆 📢
BBM 2025 will be held on June 9-10th at the Harvard Science Center feat. the one and only Dr. Petra Levin as keynote! We can't wait to see you there.
Registration and scholarship applications open next week.
01.02.2025 15:05
👍 50
🔁 30
💬 1
📌 3
PNAS
Proceedings of the National Academy of Sciences (PNAS), a peer reviewed journal of the National Academy of Sciences (NAS) - an authoritative source of high-impact, original research that broadly spans...
Angelika Gründling’s work from her sabbatical in the Bernhardt-Rudner labs is out. She came with the goal of finding the missing G+ phosphatidylglycerol phosphate phosphatase and found it with time to spare. I can also highly recommend her as a bay mate! www.pnas.org/doi/10.1073/...
28.01.2025 15:10
👍 9
🔁 3
💬 0
📌 0

HARIBO bacter for happy holidays
20.12.2024 15:38
👍 5
🔁 2
💬 0
📌 0