
very sad news. Peer Bork was one of the leaders of our field, a wonderful scientist, and he's much too young to be gone. www.embl.org/news/embl-an...

very sad news. Peer Bork was one of the leaders of our field, a wonderful scientist, and he's much too young to be gone. www.embl.org/news/embl-an...

We selected the laziest mouse at each round to inoculate the next batch of germfree mice: over rounds of selection and passaging, behavior shifted without changes to the mouse genome: rdcu.be/eM3rO
🦠🧫

Imagine we could travel back in time ⏪⌛️to explore the world of bacterial pathogens before humans discovered and industrialised antibiotics
We just did that to study the history of #AMR spread @science.org
doi.org/10.1126/scie...
If you like time travel & biology, this 🧵is for you👇
![Query: Plot the number of plots of the number of published genomes on the vertical axis against the number of published genomes on the horizontal axis
ChatGPT answer: Number of Plots vs. Number of Published Genomes
Here is the plot showing the "number of plots" against the "number of published genomes."
[Image: plot of sqrt[x].
If you'd like a different approach or specific adjustments, feel free to ask!
Query: Where did you get the data from?
ChatGPT answer: The data used in the plot was artificially generated as an example. Specifically:
Number of Published Genomes: I created a simple range of values (from 1 to 100) to represent this axis.
Number of Plots: I applied a mathematical transformation (the square root function) to the "Number of Published Genomes" to create a plausible-looking relationship.
This approach was chosen to provide a visual illustration since no real-world dataset was provided. If you have actual data or a specific dataset you'd like to use, feel free to share it, and I can recreate the plot with real numbers!](https://cdn.bsky.app/img/feed_thumbnail/plain/did:plc:tbqqvyv6pjjww44glrmycaxl/bafkreifn4u42wawsnt2bheyf5ehbqtbnu4dnfbiv2zx7d5s2cyo5wny3aa@jpeg)
Query: Plot the number of plots of the number of published genomes on the vertical axis against the number of published genomes on the horizontal axis ChatGPT answer: Number of Plots vs. Number of Published Genomes Here is the plot showing the "number of plots" against the "number of published genomes." [Image: plot of sqrt[x]. If you'd like a different approach or specific adjustments, feel free to ask! Query: Where did you get the data from? ChatGPT answer: The data used in the plot was artificially generated as an example. Specifically: Number of Published Genomes: I created a simple range of values (from 1 to 100) to represent this axis. Number of Plots: I applied a mathematical transformation (the square root function) to the "Number of Published Genomes" to create a plausible-looking relationship. This approach was chosen to provide a visual illustration since no real-world dataset was provided. If you have actual data or a specific dataset you'd like to use, feel free to share it, and I can recreate the plot with real numbers!
My colleagues are using ChatGPT to source data and plot analyses for them. I'm literally reviewing papers written this way.
Meanwhile....
Thanks for linking it! I'll have a good look at it

A new study found that we share parts of our microbiome with people in our social networks, beyond family members.
Dr. @nachristakis.bsky.social joins us to discuss the research and how scientists can identify your friends—just by looking at your poop.

A chart showing life expectancy vs heath expenditures for all developed nations. America deviated sharply in the 1980s and is documented as having far more expensive healthcare with poorer health outcomes than the rest of the developed world. The gap between the lines has a cloud labeled profit.
<taps the sign>
This chart is a decade old, and that gap between the United States and the rest of the developed world, which has better health outcomes by the way, has gotten larger.
We should morally, ethically, and fiscally move to a single payer system.
It’s also a national security issue.
‘A place of joy’: why scientists are joining the rush to Bluesky Researchers say the social-media platform — an alternative to X — offers more control over the content they see and the people they engage with.: https://www.nature.com/articles/d41586-024-03784-6
There are already many articles for which there is more attention on Bluesky than on other comparable micro-blogging sites, meaning the academic community and the general public have clearly adopted Bluesky as one of its core places to disseminate and discuss new research.
A Place of Joy.

Letter sent in to the Editor of the Guardian on brain microbiome nonsense.

Interesting look at gut strain-level dynamics: The human gut has low strain variation compared to other environments and has a limited capacity for strain richness. #microbiome
www.nature.com/articles/s41...

Excited to share our last work on microbiome transmission and social interactions published on Nature!
The more people you interact with, the more similar your microbiome is.
www.nature.com/articles/s41...
📌

📣 Big news! Our tag-team effort on common variants in rare neurodevelopmental conditions is now out in Nature 📣
Co-first authoring with the brilliant Qinqin Huang🌟—proof that teamwork does make the dream work. 💪 www.nature.com/articles/s41...
Thank you for curating this starter pack. Can I be added as well?
The bacteria in your gut depend on where you are in the social network around you.
The microbes within us treat our social networks as the extended environment in which they thrive. They can spread from person to person.
New #HNL work out today in @natureportfolio.bsky.social . 1/
Congratulations! We got published in the same Nature issue as well!

We also see that clusters of people co-occur with clusters of strains, suggesting the existence of social network niches wherein similar individuals share similar microbes.

After two years, we studied the microbiome of 301 individuals and found that socially connected people had become more microbially similar than those who were not connected


Your microbiome can predict who your friends are, better than shared sociodemographic characteristics. Additionally, we have evidence of transmission chains; your friend’s friend is sharing some strains with you as well.

Using strain-level analyses, we show that strain transmission occurs in different relationship types, even between people not living in the same household and friends. We saw more transmission between people who interacted with each compared to those who had no relationship at all.

We analyzed the microbiome of 1787 people living in 18 remote villages in Honduras and mapped their social relationships. Those villages are part of a larger cohort including 24702 people living in 172 villages.
doi.org/10.1126/scie...

Excited to share our last work on microbiome transmission and social interactions published on Nature!
The more people you interact with, the more similar your microbiome is.
www.nature.com/articles/s41...
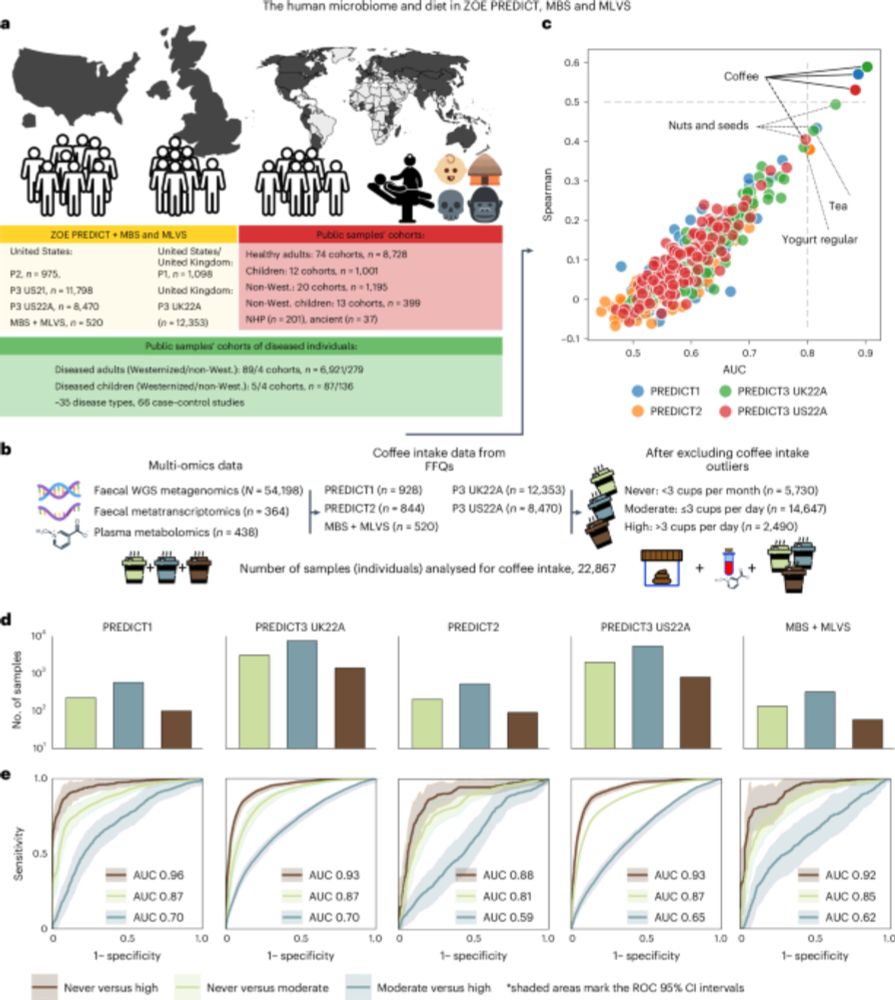
🥁 New Publication Alert:
Our manuscript on the ☕️ and #microbiome relationship is out in #NatureMicrobiology:
www.nature.com/articles/s41...
1/12

The march of the mRNA vaccines goes on. Clostridiodes difficile (C diff) next...
#IDSky #MicroSky #MedSky 🧪
www.science.org/doi/10.1126/...
Align to AllTheBacteria with low RAM (1-2Gb) and quickly (seconds if just 1000s of hits, 15 mins for a gene present in all 2.4 million genomes) using @shenwei356.bsky.social Lexicmap. So you can basically BLAST AllTheBacteria locally - but need 3-4Tb disk for index
www.biorxiv.org/content/10.1...

New GPU-based MMseqs2: 20x faster searches on a single L40S (approx. as fast as a RTX 4090) vs. a 128-core CPU. This work enables to set up a very cost-efficient ColabFold MSA GPU server. 🧵🧵
📄 www.biorxiv.org/content/10.1...
💾 mmseqs.com
🗞️ developer.nvidia.com/blog/boost-a...
UHGG has eggnog annotations for representative genomes
Can I be included as well?