
(1/n) Excited to present the latest work from the de Wit lab, where we identify and characterise loop extrusion-mediated fountains in mammalian genomes using acute depletion of 3D genome regulators: doi.org/10.1093/nar/.... A Bluetorial🧵:

(1/n) Excited to present the latest work from the de Wit lab, where we identify and characterise loop extrusion-mediated fountains in mammalian genomes using acute depletion of 3D genome regulators: doi.org/10.1093/nar/.... A Bluetorial🧵:
The review also highlights exciting applications of SMT to nuclear factors and related findings, along with a perspective of future directions, including complementary genome-wide measurements of kinetic parameters.
I hope this review further sparks interest in and the application of live-cell SMT!

and a comprehensive discussion of kinetic parameters accessible by SMT.

a discussion of temporal excitation patterns that facilitate measuring kinetic properties,

an overview of SMT microscopy techniques,

In the review, I give an overview of important steps along the implementation, execution, and analysis of single-molecule tracking in living cells and multicellular organisms. This includes a summary of labeling approaches,

The review on live-cell SMT that I contribute to the jmolbiol.bsky.social special issue ‚Imaging of the central dogma‘ is now online as pre-proof: www.sciencedirect.com/science/arti...

This is a piece that I and @karsten-rippe.bsky.social discussing a lot, and a topic that is very close to my heart. The editors @naturerevgenet.bsky.social gave us the stage to do so, and the final version of our review is now available under this link: rdcu.be/erP1u
A short thread follows 1/n
So beautifully played :-) All the best for the final!
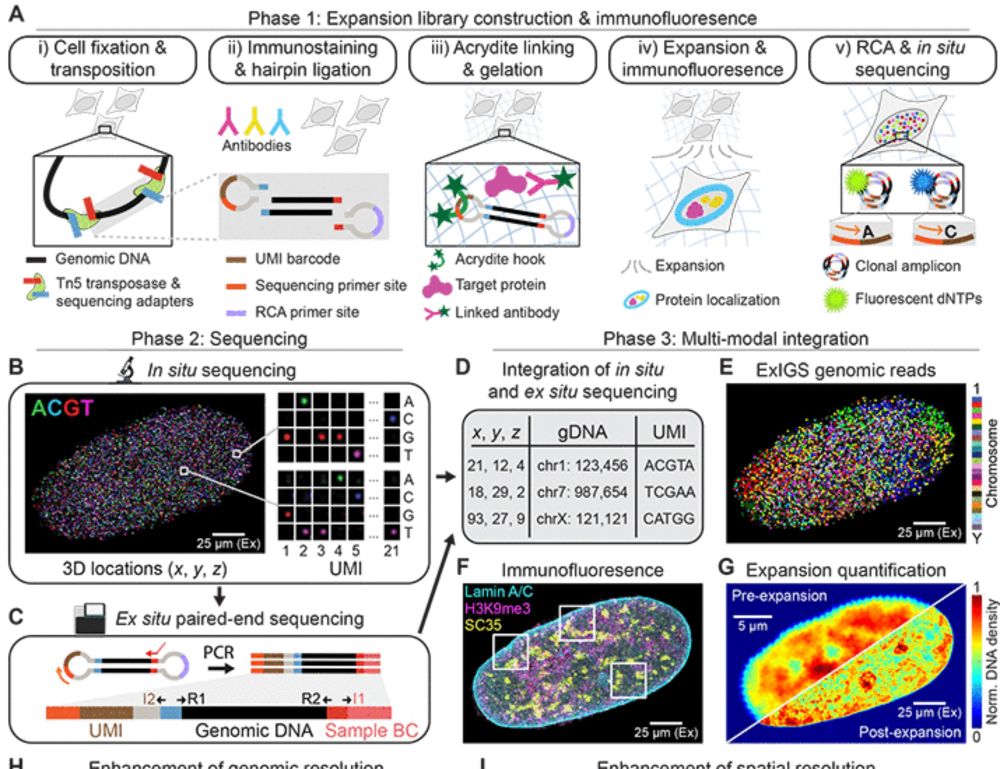
@science.org Expansion in situ genome sequencing links nuclear abnormalities to aberrant chromatin regulation | Science www.science.org/doi/10.1126/...
@jbuenrostro.bsky.social @eboyden3.bsky.social + al @broadinstitute.org @harvard.edu
#ExIGS #Chromatin #Lamin #Progeria #Aging #ExpansionMicroscopy

Great to see this published @genesdev.bsky.social. Evidence that enhancers can activate a bystander gene in an adjacent TAD, and that cohesin facilitates this. @uoe-igc.bsky.social.
pubmed.ncbi.nlm.nih.gov/40436628/


1/
Super excited to share my first preprint from @andersshansen.bsky.social Lab! We used MINFLUX to track chromatin (H2B-Halo and Fbn2 locus) at an unprecedented 200 μs, then combined it with SPT to span μs-minutes (H2B) or SPT & Super-Res Live-Cell Imaging (SRLCI) to span μs-hours (Fbn2)
📣 New preprint from the lab – Massive work entirely driven by super-talented @simsakong.bsky.social , jointly betw. @beatfierz.bsky.social lab and mine – combining in vitro and in vivo TF single molecule imaging
www.biorxiv.org/content/10.1...
Fun introduction!
Intriguing mechanism to balance production and degradation rates of proteins
Fantastic! Your playing is really great!

Program is out for our first EMBL #singlemolecule gene regulation meeting! Check out the dedicated panel sessions on single molecule genomics, single molecule imaging and theory where we would like to brainstorm the future of the field:
Registration is open! 👇
www.embl.org/about/info/c...

Search gets faster, binding longer, bound fraction of cohesin and CTCF goes up, especially at and after ZGA

Single molecules, in living zebrafish embryos, during hour-long period of development: Jonas' work on binding kinetics of cohesin and CTCF around ZGA now online. Great collab with labs of @lucagiorgetti.bsky.social @akispapantonis.bsky.social
doi.org/10.1038/s414...
#Cellpose 3 paper now out. Not all images are perfect. Restore your images with Cellpose3 to get better segmentations, w/ @marius10p.bsky.social www.nature.com/articles/s41...

Our new review on how the #chromatin domain is formed in the cell is now available @Curr Opin Struct Biol.📄✨
We critically discuss the domain formation mechanism from a physical perspective, including #phase-separation and #condensation.
📥 Free-download link:
authors.elsevier.com/a/1keGn,LqAr...
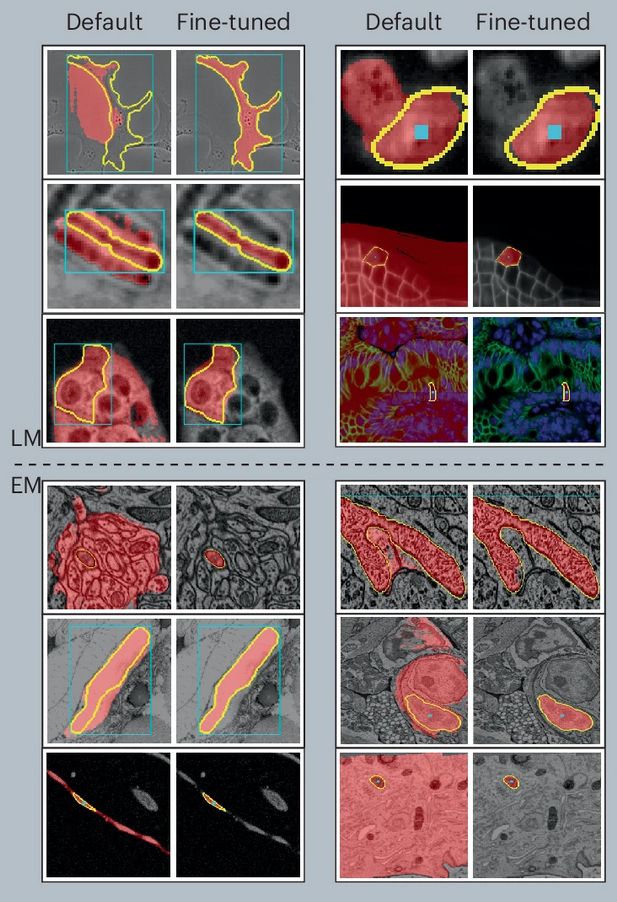
Improvements in LM (top) and EM (bottom) of our micro-sam model (finetuned) compared to the default SAM model.
After a long journey, Segment Anything for Microscopy is now published in Nature Methods! We significantly improve SAM for interactive and automatic segmentation in light and electron microscopy and build a user-friendly tool.
www.nature.com/articles/s41...
Fantastic work, @armelletollenaere.bsky.social , @davidsuter.bsky.social and co-workers! Happy we could contribute.
Big thanks to all people involved @armelletollenaere.bsky.social , Enes Ugur, Susanna Dalla-Longa, Cédric Deluz, Devin Assenheimer, @gebhardtlab.bsky.social , and Heinrich Leonhardt
And thanks ChatGPT for haikuing the abstract
1/n Excited to share our new preprint! Using live imaging in Drosophila embryos, we show that RNA Pol II clusters switch from sites of initiation to elongation during zygotic genome activation and that they are stably associated with an active gene: www.biorxiv.org/content/10.1...
Great work and very useful developments!
Measuring morphogen transport over multiple spatial scales in live zebrafish embryos https://www.biorxiv.org/content/10.1101/2025.02.06.636788v1
After not having been on Twitter for a year, I somehow missed that everyone had migrated already.
Thank you! Happy this finally made it.
