Absolutely! He was always at Wednesday journal clubs while I was a post-doc at the NMNH and, in addition to enjoying a cookie, had lots of opinions and stories to share on any topic. A real inspiration!
Absolutely! He was always at Wednesday journal clubs while I was a post-doc at the NMNH and, in addition to enjoying a cookie, had lots of opinions and stories to share on any topic. A real inspiration!

A legend has left us. RiP Brian. Thank you for creating the soundtrack to my formative years. www.nytimes.com/2025/06/11/a...

I'm sure you've all seen this already but it's too good not to share. www.nytimes.com/2025/04/21/o...
I've just been listening to RATM's first album on loop
Hello graduate students! Volunteer travel award applications are due March 21, 2025. In return for working two ~4h shifts, graduate student members of one or more of our societies can receive a rebate equal to 100% of the regular early student member registration fee.
The only small silver lining - quitting Amazon has already been good for my bank balance, and quitting twitter and facebook has been good for my brain.

A grayscale illustration of a hyena skull on turquoise background, overlaid by letters and a network of circles that represent the complex interconnections of skull bones produced by macroevolutionary change.
New, NSF-supported research: Emergent network properties link phenotypic modules to ecomorphological divergence in carnivoran mammals doi.org/10.1016/j.is...
Also, check out the beautiful original artwork made by one of the co-authors for this study:
Pet Sounds is the perfect soundtrack to anything!
oh no, im here teaching while all the students get to have fun in Georgia
tell me about it. I'm in denial about our quarter starting today

A reminder that we have two open PhD projects on Bayesian phylogenetics in the lab:
One with the new TREE Doctoral Landscape Awards: www.trees-dla.ac.uk/projects/int... (deadline 20th Jan)
And another with CSC: www.findaphd.com/phds/project... (deadline 29th Jan)
#SICB2025 isn't over yet, right. #UChicago grad student Menna Jones will be talking about her work exploring modes of habitat dependent functional trait evolution in frogs on Tues morning at 8am in International Salon 5 🐸
I'm so excited to share our range-wide paper on #snowleopard phylogeography & population structure! Representing my PhD work & using the largest sample size & geographic coverage to date, our work fills important knowledge gaps on snow leopard gene flow: rdcu.be/dWNKj 🦊 🧬🧪
Tomorrow means Monday at 10.45am btw
I'm not at #SICB2025, but lab members are. Check out Mags Mercado, talking about "Intraspecific variability in vertebral formulae in marsupials and monotremes" in A601 at 10.45am tomorrow! Mags is defending her proposal this spring and is excited to get feedback!
You're welcome! I'm pleased to know I'm not the only one still using it!
If you didn't solve this yet, try downloading the centos version - that one works just fine on ubuntu 24.04
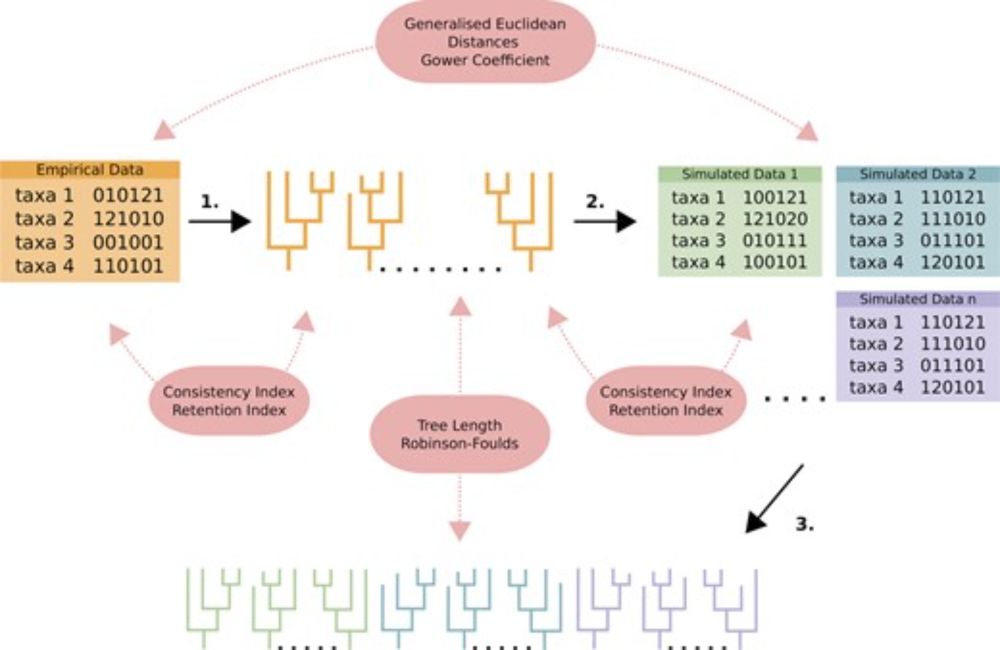
So excited to see this paper finally published! 🥳 This was the main focus of my PhD and demonstrates the application of model adequacy for models of morphological evolution. 1/n
academic.oup.com/sysbio/advan...
Huge congratulations to @pollardmd.bsky.social on the publication of his first 1st author paper!
Our paper with @sorrywm.bsky.social is out at @genomeresearch.bsky.social
I'll do a thread of the main results. 🧪🧬 #evolution #evosky
1/n
www.genome.org/cgi/doi/10.1...
Thinking of starting a PhD on Bayesian phylogenetics next October 2025 in London? 🎓
You may want to check the TREES project led by the dos Reis Lab on integrating morphology and genomes for timetree inference! 🦴🧬💻Project supervisors: @mariodosreis.bsky.social & Ziheng Yang. More info in link 🔽
You're already there, James. You think I'd forget about the king of the pinnipeds?
This is a first step, but one we think could be fruitful. Feedback is very welcome. David isnt on here yet, but please email us if you have thoughts or suggestions!
We've only investigated its performance for the ultrametric case so far but it should work well for non-ultrametric cases too. However, we still need to implement an imputation algorithm appropriate to such data. Scalability beyond 1K tips remains a challenge but David has ideas....
The result is an approach that takes a set of time scaled trees with partially or incompletely overlapping leaf sets as input, does some stuff, and then samples from the posterior distribution of chronograms over the entire leaf set. It seems to work reasonably well ..
David did this by combining and tweaking two distinct supertrees approaches - the least squares-based average consensus approach of Levasseur and Lapointe, and the probabilistic exponential error approach of Steel and Rodrigo, and recasting the problem as one of sampling rather than optimizing
that second part is key; while molecular phylogenies have increased massively in size over the past decade, the upper limits on morphological dataset sizes remain relatively low (~200 tips with a couple outliers). in other words, mega paleophylogenies are going to be supertrees
This is all great but we had 2 questions - first, can we come up with a bayesian supertree approach that can sample directly from the posterior distribution of chronograms, and second, can we ensure this approach can be extended to non-ultrametric cases
In the past few years, we've seen larger and larger "megaphylogenies" of extant clades, most of which either "patch" time-scaled subtrees onto a time-scaled "backbone" or non-parametrically smooth rates to time scale a single ML tree