e-DNA surveys could improve marine biosecurity – Australian Antarctic Program (News 2026)
Antarctic scientists have trialed a DNA ‘barcoding’ technique that could improve biosecurity measures that help protect polar ecosystems from invasive marine species.
Another belated "new paper". We used passive #eDNA sampling to evaluate the risk of biofouling species being introduced to Macquarie Island (and other Antarctic / subantarctic locations) by ships.
Story from AAD: www.antarctica.gov.au/news/2026/ed...
and link to paper: doi.org/10.1016/j.sc...
🧬🧪
06.03.2026 02:17
👍 14
🔁 3
💬 0
📌 0


Another day, another estuary. Sampling #eDNA to better understand biodiversity in Tilba Tilba Lake
An important aspect our work is communication with local stakeholders, such as the farmers around this estuary
Visits from cute echidnas is an added bonus
#eDNAinFocus @sednasociety.bsky.social
06.03.2026 03:26
👍 7
🔁 2
💬 0
📌 0
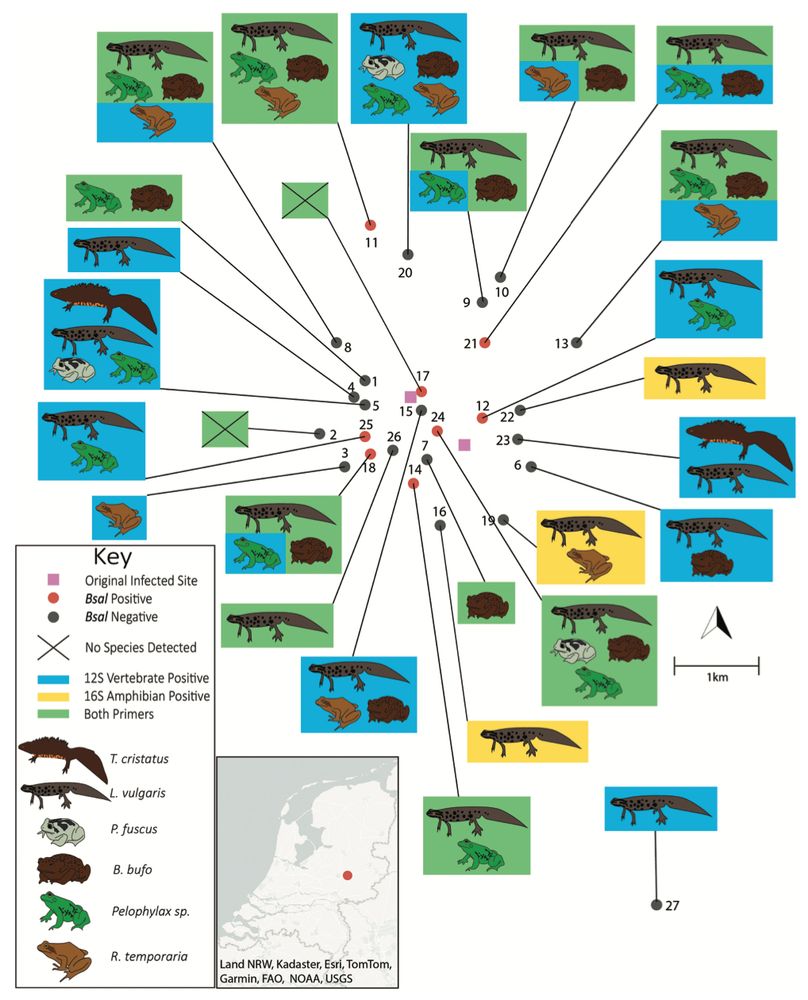
New paper out from our crew on using #eDNA to detect the pathogenic chytrid fungus (Bsal) and characterise the surrounding amphibian community from pond samples in the Netherlands. Small number of samples but highlights the potential of eDNA for routine monitoring of both. bioone.org/journals/jou...
24.06.2025 10:47
👍 21
🔁 10
💬 0
📌 0
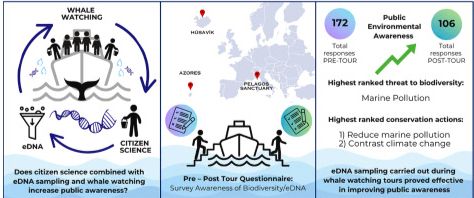
New @ewhale-dna.bsky.social publication led by Eleonora Barbaccia & published in Ocean & Coastal Management! 🐋💙
#WhaleWatching, #citizenScience, and #eDNA sampling can foster public awareness and engagement in #marine #biodiversity #conservation
doi.org/10.1016/j.oc...
20.06.2025 14:00
👍 9
🔁 5
💬 0
📌 1

#newepisode just dropped!
Environmental #DNA or #eDNA is the study of genetic material in the #environment.
How can we apply it to the #deepsea? Revealing the hidden majority, changes through time and community structure, the possibilities seem endless.
open.spotify.com/episode/2XQv...
06.06.2025 22:06
👍 13
🔁 6
💬 1
📌 0
1/4 A major scientific breakthrough happened during the last Vendée Globe! ⛵️ Fabrice Amedeo collected environmental DNA (eDNA) for 114 days, documenting marine life across 3 oceans, 100 degrees of latitude, and 360 degrees of longitude for the first time. #eDNA #OceanBiodiversity
07.06.2025 17:03
👍 4
🔁 1
💬 1
📌 0

Robotic environmental DNA bio-surveillance of freshwater health - Scientific Reports
Scientific Reports - Robotic environmental DNA bio-surveillance of freshwater health
Autonomous water sampling tech like MBARI's ESP can revolutionize long-term river monitoring by automating #eDNA collection offering cost-effective, high-frequency detection of pathogens & invasive species.
Read more: bit.ly/45YBE0o
#Innovation #Tech #Monitoring #Science
06.06.2025 07:38
👍 5
🔁 2
💬 0
📌 0
Proud of this new publication demonstrating #eDNA as a useful accessory to invasive species management in practice. In this case, we observed eDNA declines coinciding with community-led apple snail removal from an Austin, Texas pond. www.reabic.net/journals/mbi...
04.06.2025 21:09
👍 19
🔁 7
💬 2
📌 0

New #sharkscience led by Jorge Valerio-Vargas in @esr-ir.bsky.social
Using #eDNA to find remnant sawfish populations in Costa Rica to aid in prioritizing conservation efforts.
@saveourseas.bsky.social @iucnshark.bsky.social
www.int-res.com/abstracts/es...
06.06.2025 04:32
👍 20
🔁 4
💬 0
📌 0

🌿 #InternationalDayBiologicalDiversity
At Genome BC, we’re investing in #genomics research that helps protect and preserve biodiversity — from tracking endangered species with #eDNA to understanding how microbes support ecosystem health.
22.05.2025 22:08
👍 3
🔁 1
💬 1
📌 0

eDNA Surveillance
📢 JUST ANNOUNCED: Genome Canada invests in #eDNA surveillance to safeguard health, environmental protection and Canadian industries.
📲 bit.ly/4kk2buX
14.05.2025 16:42
👍 14
🔁 5
💬 0
📌 0
Our review is finally here! We include all the terrestrial and semi-aquatic mammal-focused #eDNA and #iDNA studies available up to Jan 2025! It's amazing to see how far this field has come 🧬🦦
05.05.2025 11:43
👍 17
🔁 10
💬 0
📌 1
OceanOmics - Registry of Open Data on AWS
Our OceanOmics AWS Open Data repository is now live!
registry.opendata.aws/oceanomics/
It holds all of our #eDNA data and all of our curated marine vertebrate genome assemblies.
There's so far only a single assembly in there - I'm currently gathering all the eDNA raw reads for upload.
02.05.2025 02:12
👍 8
🔁 2
💬 1
📌 1

Junior faculty job alert in Germany - Marine Evolutionary Genomics @geomarkiel.bsky.social / University of Kiel www.geomar.de/en/karriere/... Marine study system of your choice (no microbes, sorry…), excellent infrastructure and ship access, moderate teaching requirements, closing date 6 June
26.04.2025 15:10
👍 74
🔁 92
💬 1
📌 4

Rediscovering Lost Amphibians: Using eDNA to Detect Critically Endangered Frogs in Área de Conservación Guanacaste, Costa Rica
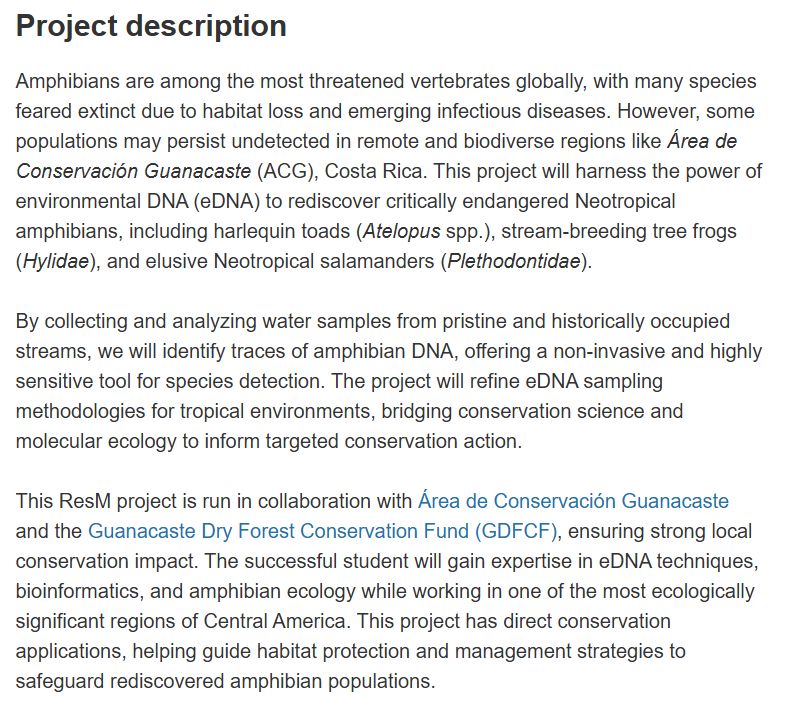

Project description
Amphibians are among the most threatened vertebrates globally, with many species feared extinct due to habitat loss and emerging infectious diseases. However, some populations may persist undetected in remote and biodiverse regions like Área de Conservación Guanacaste (ACG), Costa Rica. This project will harness the power of environmental DNA (eDNA) to rediscover critically endangered Neotropical amphibians, including harlequin toads (Atelopus spp.), stream-breeding tree frogs (Hylidae), and elusive Neotropical salamanders (Plethodontidae).
By collecting and analyzing water samples from pristine and historically occupied streams, we will identify traces of amphibian DNA, offering a non-invasive and highly sensitive tool for species detection. The project will refine eDNA sampling methodologies for tropical environments, bridging conservation science and molecular ecology to inform targeted conservation action.
This ResM project is run in collaboration with Área de Conservación Guanacaste and the Guanacaste Dry Forest Conservation Fund (GDFCF), ensuring strong local conservation impact. The successful student will gain expertise in eDNA techniques, bioinformatics, and amphibian ecology while working in one of the most ecologically significant regions of Central America. This project has direct conservation applications, helping guide habitat protection and management strategies to safeguard rediscovered amphibian populations.

Eligibility
Applicants should have a first or upper second-class honours degree in a relevant subject such as biological sciences, molecular ecology, conservation biology, or a related discipline. A relevant Master’s qualification is desirable but not essential.
This project is primarily laboratory-based, and applicants should have experience or a strong interest in molecular techniques, particularly environmental DNA (eDNA) analysis, PCR, and bioinformatics. Experience with DNA extraction, qPCR, metabarcoding, and sequence data analysis is advantageous. A background in amphibian ecology, conservation genetics, or freshwater ecology would also be beneficial.
The ideal candidate will be highly motivated, detail-oriented, and comfortable working with environmental samples in a molecular laboratory setting. Strong analytical skills and the ability to work independently while collaborating with conservation partners are essential.
If your first language is not English, you must meet the minimum English language requirements for the programme: an IELTS Academic score of 6.5 (with no less than 5.5 in each component test area) or an equivalent qualification.
The project is supported for 1.5 years and covers full tuition fees only.
If you wish to discuss this project further informally, please contact Dr Robert Puschendorf by email: robert.puschendorf@plymouth.ac.uk.
Please see our
apply for a postgraduate research programme
page for a list of supporting documents to upload with your application.
🧪
Student Opportunity!!
Interested in using #eDNA to rediscovering lost #amphibian species in Área de Conservación Guanacaste, Costa Rica?
Check out this opportunity led by @rpuschen.bsky.social and join us working with #ACG and @gdfcf.bsky.social
🔗 Info: www.plymouth.ac.uk/study/resear...
🧪
16.04.2025 14:24
👍 20
🔁 15
💬 0
📌 0
🚨 NEW PAPER 🚨
Real-Time PCR Assay and Environmental DNA Workflow for Detecting Irukandji Jellyfish, Malo bella (Cubozoa)
Led by Jess Strickland, this study presents a new #eDNA assay for detecting M. bella - a small but nasty Irukandji jellyfish found along WA’s Ningaloo Reef
tinyurl.com/3yvvnxzc
15.04.2025 01:48
👍 31
🔁 10
💬 2
📌 1
Cool new aerial #eDNA preprint which shows the potential to contribute to large scale biodiversity monitoring. 🧪🧬🌍🖥️
13.04.2025 08:27
👍 21
🔁 3
💬 0
📌 0

📣Excited to share that our new #paper is out! Using #eDNA #metabarcoding to detect #elasmobranchs in #OWFs in the North Sea🦈🦈
www.sciencedirect.com/science/arti...
@reindertn.bsky.social
@nanoporetech.com
10.04.2025 05:22
👍 9
🔁 6
💬 0
📌 0
Awesome to see our colleagues’ work rearing at the Sunflower lab highlighted!
We’re working with them and others echinodermologists across the West Coast to design an #eDNA assay to help find the last CA sunflower stars and enhance future monitoring efforts!
11.04.2025 15:37
👍 40
🔁 12
💬 4
📌 0