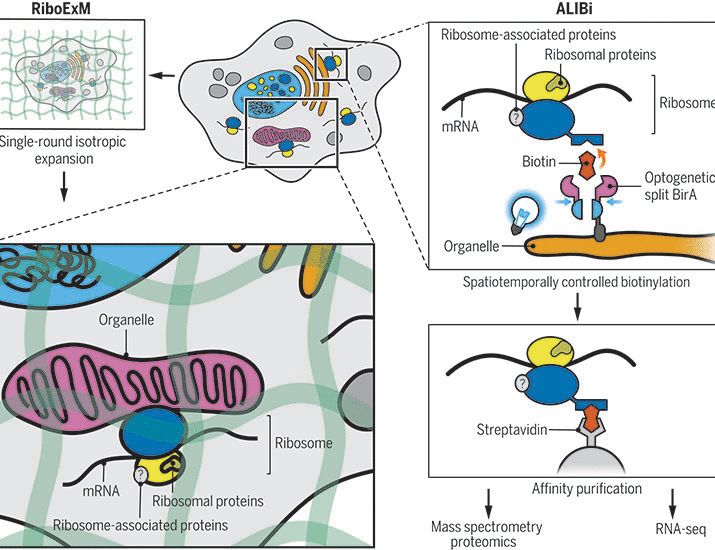
Overall, tRIBO-seq provides a simple way to profile the active tRNAome, capturing abundance, modifications and fragmentation from actively translating ribosomes.
This project was co-led with Hasan Yilmaz, in collaboration between the Novoa Lab, IMMAGINA Biotech and the Soares Lab.
💃
04.03.2026 09:25
👍 4
🔁 0
💬 0
📌 0
Under basal conditions, the ribosome-associated tRNA pool largely matches the total cellular tRNA pool.
But under stress, the picture changes...
04.03.2026 09:25
👍 4
🔁 0
💬 1
📌 0
Using @nanoporetech.com sequencing allows us to capture multiple layers of tRNA biology in a single experiment:
• tRNA abundance
• modification signatures
• fragmentation patterns
All directly from ribosome-associated tRNAs.
04.03.2026 09:25
👍 2
🔁 0
💬 1
📌 0

tRIBO-seq builds on RiboLace technology from to pull down active ribosomes.
Using ~15× less starting material, we recover comparable ribosome-associated tRNAs to polysome purification - with a much faster protocol.
Importantly, the recovered tRNAs correlate well with polysome-derived samples.
04.03.2026 09:25
👍 2
🔁 0
💬 1
📌 0

To address this, we developed tRIBO-seq alongside @immagina.bsky.social.
The idea is simple:
• capture actively translating ribosomes
• isolate their associated native tRNAs
• sequence them using nanopore
This provides a direct view of the ribosome-associated tRNA population (ribo-tRNAs).
04.03.2026 09:25
👍 3
🔁 0
💬 1
📌 0
tRNAs deliver amino acids to ribosomes according to mRNA codons.
But most studies measure total cellular tRNAs, not the tRNAs actually engaged with translating ribosomes.
This makes it hard to understand the role of the active tRNA pool in translation 🧐.
04.03.2026 09:25
👍 2
🔁 0
💬 1
📌 0

New preprint from
@novoalab.bsky.social !
Which tRNAs are used by ribosomes during translation?
We introduce tRIBO-seq, a nanopore method to sequence ribosome-associated tRNAs and track how the active tRNA pool changes across stress conditions.
www.biorxiv.org/content/10.6...
Thread 👇
04.03.2026 09:25
👍 34
🔁 13
💬 2
📌 1
Using @nanoporetech.com sequencing allows us to capture multiple layers of tRNA biology in a single experiment:
• tRNA abundance
• modification signatures
• fragmentation patterns
All directly from ribosome-associated tRNAs.
04.03.2026 09:15
👍 0
🔁 0
💬 0
📌 0

tRIBO-seq builds on RiboLace technology to pull down active ribosomes.
Using ~15× less starting material, we recover comparable ribosome-associated tRNAs to polysome purification - with a much faster protocol.
Importantly, the recovered tRNAs correlate well with polysome-derived samples.
04.03.2026 09:15
👍 0
🔁 0
💬 1
📌 0

To address this, we developed tRIBO-seq alongside @immagina.bsky.social
The idea is simple:
• capture actively translating ribosomes
• isolate their associated native tRNAs
• sequence them using nanopore
This provides a direct view of the ribosome-associated tRNA population (ribo-tRNAs).
04.03.2026 09:15
👍 0
🔁 0
💬 1
📌 0
tRNAs deliver amino acids to ribosomes according to mRNA codons.
But most studies measure total cellular tRNAs, not the tRNAs actually engaged with translating ribosomes.
This makes it hard to understand the role of the active tRNA pool in translation.
04.03.2026 09:15
👍 0
🔁 0
💬 1
📌 0
tRNAs deliver amino acids to ribosomes according to mRNA codons.
But most studies measure total cellular tRNAs, not the tRNAs actually engaged with translating ribosomes.
This makes it hard to understand the role of the active tRNA pool in translation.
04.03.2026 08:51
👍 0
🔁 0
💬 0
📌 0
The lab of Single cell ecology has just opened its doors at the University of Glasgow, School of Biodiversity, One Health and Veterinary Medicine!
Join us, collaborate or learn about what we do:
mancinilab.github.io/index.html
And check out our most recent output:
www.biorxiv.org/content/10.1...
02.12.2025 10:33
👍 2
🔁 1
💬 0
📌 1

It was our pleasure to host @james-bryson.bsky.social (BRIC, Copenhagen) at @crg.eu through the EU-LIFE Postdoctoral Exchange Program!
Big thanks to James for presenting his work on advanced CRISPR tools for epitranscriptomic research, and to @eu-life.bsky.social for this opportunity!🧬✈️
#EULIFE #CRG
10.11.2025 15:59
👍 9
🔁 5
💬 0
📌 0
@embl.org is undoubtedly one of the best place in the world to start your lab! The GB unit is collegial, supportive, collaborative and the science is just outstanding. Apply! DM me if you have questions.
30.07.2025 14:01
👍 14
🔁 7
💬 1
📌 0



Last month, the @novoalab.bsky.social attended the Premis Nacionals de Recerca ceremony to celebrate @evamarianovoa.bsky.social receiving the National Young Researcher Award! 🎉👏
Huge congratulations again, Eva - an incredible and well-deserved recognition! 🧬🏆
22.07.2025 15:50
👍 7
🔁 2
💬 0
📌 0
Sciencing in the summer
11.07.2025 16:44
👍 1
🔁 0
💬 0
📌 0
Would you pay $150 per article to post your preprint on @bioRxiv @aRxiv @medRxiv @chemRxiv etc.? Currently it’s free but a small cost could help offset running costs and also work as a barrier against paper mills and AIs just posting all the time and filling these sites up.
09.07.2025 18:03
👍 88
🔁 19
💬 25
📌 5
Decoding cnidarian cell type gene regulation
Animal cell types are defined by differential access to genomic information, a process orchestrated by the combinatorial activity of transcription factors that bind to cis -regulatory elements (CREs) to control gene expression. However, the regulatory logic and specific gene networks that define cell identities remain poorly resolved across the animal tree of life. As early-branching metazoans, cnidarians can offer insights into the early evolution of cell type-specific genome regulation. Here, we profiled chromatin accessibility in 60,000 cells from whole adults and gastrula-stage embryos of the sea anemone Nematostella vectensis. We identified 112,728 CREs and quantified their activity across cell types, revealing pervasive combinatorial enhancer usage and distinct promoter architectures. To decode the underlying regulatory grammar, we trained sequence-based models predicting CRE accessibility and used these models to infer ontogenetic relationships among cell types. By integrating sequence motifs, transcription factor expression, and CRE accessibility, we systematically reconstructed the gene regulatory networks that define cnidarian cell types. Our results reveal the regulatory complexity underlying cell differentiation in a morphologically simple animal and highlight conserved principles in animal gene regulation. This work provides a foundation for comparative regulatory genomics to understand the evolutionary emergence of animal cell type diversity. ### Competing Interest Statement The authors have declared no competing interest. European Research Council, https://ror.org/0472cxd90, ERC-StG 851647 Ministerio de Ciencia e Innovación, https://ror.org/05r0vyz12, PID2021-124757NB-I00, FPI Severo Ochoa PhD fellowship European Union, https://ror.org/019w4f821, Marie Skłodowska-Curie INTREPiD co-fund agreement 75442, Marie Skłodowska-Curie grant agreement 101031767
I am very happy to have posted my first bioRxiv preprint. A long time in the making - and still adding a few final touches to it - but we're excited to finally have it out there in the wild:
www.biorxiv.org/content/10.1...
Read below for a few highlights...
06.07.2025 18:14
👍 59
🔁 24
💬 1
📌 2

Endinsa’t en una vivència sensorial única que connecta ciència i art immersiu. “SEEING: Un ull i una caixa” arriba del 5 al 14 de juny al Centre Cívic Convent de Sant Agustí (Carrer del Comerç 36, Barcelona). www.crg.eu/en/event/see...
30.05.2025 11:24
👍 9
🔁 17
💬 0
📌 1
Our genome encodes > 7000 microproteins that have been previously ignored in genome annotations. How do these microproteins impact tissue function? Here, we use an in vivo single-cell CRISPR screen to systematically explore microprotein function in the mouse epidermis. 1/9
19.03.2025 15:44
👍 66
🔁 17
💬 3
📌 5
Scientific research is a strategic necessity for resilience and competitiveness in Europe!
"While the minimum basis for a thriving European Research & Innovation is a stand-alone, well budgeted #FP10, we urge the European Commission to further follow 5 key actions"
lnkd.in/d2mzN_zu
19.03.2025 10:22
👍 5
🔁 4
💬 0
📌 0
Philosophical Transactions of the Royal Society B: Biological Sciences: Vol 380, No 1921
Excited to see issue on ribosome heterogeneity and specialisation out @royalsociety.org Philosophical Transactions B. Thanks to authors & co-editors @liamfaller.bsky.social @andershlund.bsky.social Maria Barna. Wonderful cover Bulat Fatkhullin! #ribosomes royalsocietypublishing.org/toc/rstb/202...
06.03.2025 14:59
👍 14
🔁 10
💬 0
📌 2