Does anybody know of any UK-based postdocs that are microbiome / sustainable ag related which are either out or coming up soon and need someone who can run a field site like the military and has an excellent track record in bioinformatics???
Does anybody know of any UK-based postdocs that are microbiome / sustainable ag related which are either out or coming up soon and need someone who can run a field site like the military and has an excellent track record in bioinformatics???

💾 snp-dists 1.2.0
Major upgrade to this distance matrix tool - huge performance gains from multithreading (-j), max distance short circuiting (-x) and lower triangle output (-L).
#bioinformatiocs #microbiology #genomics
github.com/tseemann/snp...
This is why @wytamma.bsky.social and I have been writing SNIPPY-NG ! It will support short reads, long reads, and assemblies as inputs. You can check it out at github.com/centre-patho...
It's not ready for prime-time yet, but it's useable!

Slightly out of context but snippy too is a great variant calling tool.. @torstenseemann.bsky.social could add support for long reads? 😄
github.com/tseemann/sni...

LLMs.
They promote suicide and murder, undermine a century of educational practice, drive deskilling, harm the environment, and benefit foreign adversaries spreading disinformation.
But the federal government wants to prohibit states from regulating them.
You'd think they were guns.

Now available online: the new 2.0 version of gutSMASH, with capabilities to detect 12 new types of catabolic gene clusters relevant to gut microbiome ecology, as well as predictions of their regulation through transcription factor binding site detection. www.sciencedirect.com/science/arti...
Amazing tool. Any plans to release a singularity or docker container of this. Will be of massive help..
Sorry meant 0.1%. A lot of tinkering at db level might be needed before the illumina v4-v5 could be analysed.
Would it make sense to create otu bins till 99% at 1% increments and gather a representative sequence? The repset could be made into a strain blast db or bowtie index. No. Of mapped reads to strain db divided by total reads should give strain level relative abundance from v4 regions.
Full length 16S data is from Sanger seq or extracted from wgs denovo assembly?

Papers from research teams with a substantial number of beginners are highly disruptive and innovative
go.nature.com/4nrEvqg

Gene co-occurrence and its association with phage infectivity in bacterial pangenomes
Phil. Trans. R. Soc. B by @annecmg.bsky.social et al
with @fbaumdicker.bsky.social
royalsocietypublishing.org/doi/10.1098/...
Welcome to #bluesky team CEH shoutout to Tim from India this time!
Assistant Professor position in the Microbiome Engineering group (Prof Sahar El Aidy) at University of Amsterdam
Assistant Professor in Microbiome Imaging and Functional Analysis
werkenbij.uva.nl/en/vacancies...

Great to see our Review "The gut microbiome connects nutrition and human health" making the cover of this month's @natrevgastrohep.nature.com
@patrickveiga.bsky.social @eranelinav.bsky.social
www.nature.com/articles/s41...
Postdoc position opening in my group! Research projects: pangenomes for diverse organisms, genome evolution, biocomputing, language models. Please reach out if interested!
Hi #microbiome folks, Do you have a custom recipe that is used for anaerobic faecal sample preservation during collection and transport at room temp? Need it for samples whose fate is culturomics.
#gutmicrobiome
Hi #microbiome folks, Do you have a custom recipe that is used for anaerobic faecal sample preservation during collection and transport at room temp? Need it for samples whose fate is culturomics.
#gutmicrobiome
Keeping it behind a landing page with necessary forms and authentication is also reasonable as long as wait times are not too long imho.
There is no problem in wanting to know how many times and by whom data has been accessed. But if no personal identity info is part of the data then i see no reason why in the age of figshare, sra and other open repos intent to share is hiding behind this tactic.
Data not shared = science didnt happen.
Journals should do a test request and then send notice why such papers shouldn’t be retracted.
🧬🖥️ Postdoc positions available SOON!!
Looking for researchers with strong computational skills (programming essential) to work on microbiome & phylogenetics projects.
📅 Start on Nov 1, 2025 (flexible)
📍 Trento, Italy
DM me for details or share!
#postdocjobs #microbiome #metagenomics #phylogenetics
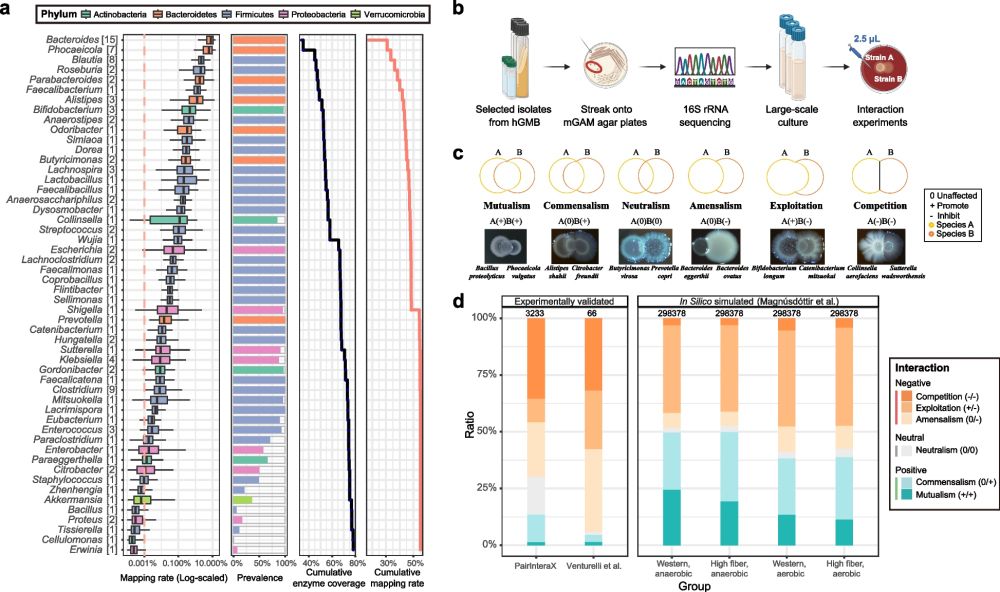
Systematic pairwise co-cultures uncover predominant negative interactions among human gut bacteria
-in BMC Microbiome
microbiomejournal.biomedcentral.com/articles/10....
If bclconvert is done then output of that too passed to multiqc for a detailed summary
Fastqc with fastp summarized using multiqc when sample numbers are very large.

I think this is a paper every microbiome scientist should read:
www.nature.com/articles/s41...

New vacancy in my team!
PhD student position on microbial genome evolution, focusing on the evolutionary principles underlying bacterial genome architecture.
Please repost and share with talented MSc students in #evobio, bioinformatics or related :)
www.uu.nl/en/organisat...
#MEvoSky #MicroSky
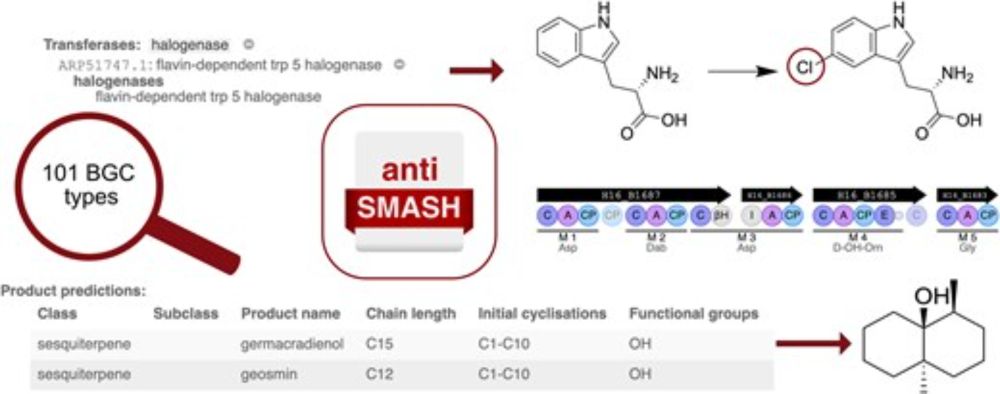
We're happy to announce that the #antiSMASH v8 paper is out in @narjournal.bsky.social: academic.oup.com/nar/advance-...
Many thanks to @kblin.bsky.social, @marnixmedema.bsky.social and many international collaborators!