Key findings:
1️⃣ Beyond Bulk: Single-nuclei distributions, not bulk averages, drive prediction.
2️⃣ Spatial Topology: GNNs reveal where these nuclei matter.
3️⃣ Chromatin Memory: This signature persists post-treatment - an intrinsic property of the resistant ecosystem.
2/3
19.12.2025 11:16
👍 2
🔁 0
💬 1
📌 0

🚨 New @biorxivpreprint! 🧬💻
Can the spatial architecture of chromatin predict chemoresponse?
We integrated FLIM imaging + ML to show that spatially organized "open" chromatin states in pre-treatment biopsies predict resistance in TNBC.
www.biorxiv.org/content/10.6...
🧵
1/3
19.12.2025 11:16
👍 4
🔁 2
💬 1
📌 0

TOMORROW is the deadline to apply for the #LakeConference on Modeling Life from Cells to Tissues!
📆 April 20-22, 2026
📍 Seattle, USA
🧠 All career stages welcomed.
Learn more and apply: https://alleninstitute.org/events/aics-lake-2026/
#StemCell #DevBio
11.12.2025 16:11
👍 0
🔁 1
💬 0
📌 0

Join us TODAY (4 pm, Room118) for the "Future of Cell Biology" session at #CellBio2025!
Hosted by the @ascbiology.bsky.social's journal Molecular Biology of the Cell (MBoC)
Hope to see you there ☺️
08.12.2025 15:24
👍 3
🔁 1
💬 0
📌 0
🤣🤣
04.12.2025 18:41
👍 1
🔁 0
💬 0
📌 0
Thanks Maria!
30.10.2025 15:51
👍 0
🔁 0
💬 0
📌 0
Somehow missed acknowledging Alon Shpigler, the first author 😱
30.10.2025 14:15
👍 0
🔁 0
💬 0
📌 0

Congratulations to Naor Kolet who led this project and thanks to co-authors Shahar Golan for his contribution and to @erinweisbart.bsky.social l for her critical insights regarding interpretability 🎉
3/3
30.10.2025 09:44
👍 0
🔁 0
💬 0
📌 0
Anomaly-based representations have several benefits:
1. Reducing batch effects
2. Improving reproducibility and mechanism of action classification
3. Complementing classical representations
4. Revealing alterations in biologically interpretable dependencies
2/3
30.10.2025 09:44
👍 0
🔁 0
💬 1
📌 0

Our paper is out ✨
doi.org/10.1016/j.ce...
We propose a self-supervised anomaly representation that encodes morphological inter-feature dependencies for high-content image-based cell phenotyping!
1/3
30.10.2025 09:44
👍 2
🔁 0
💬 3
📌 1
Join us!
10.10.2025 14:37
👍 0
🔁 0
💬 0
📌 0
Jennifer Lippincott-Schwartz, Emma Lundberg, Alex Mogilner, Maddy Parsons, @loicaroyer.bsky.social, Guillaume Salbreux, Anđela Šarić, Timm Schroeder, Hervé Turlier, @virginieuhlmann.bsky.social, Vincenzo Vitelli
Please save the date and spread the word!
3/3
10.10.2025 09:23
👍 2
🔁 0
💬 0
📌 0
Co-organized with Meghan Driscoll, @quantumjot.bsky.social and Susanne Rafelski
Speakers include: @kevin-dean.bsky.social, @ellenberglab.bsky.social, Margaret Gardel, @florianjug.bsky.social, @krishnaswamylab.bsky.social, @levayerr.bsky.social, @priscaliberali.bsky.social, ...
10.10.2025 09:23
👍 8
🔁 1
💬 1
📌 0

Excited to announce the first @cshlnews.bsky.social meeting on Cell Modeling in Space and Time, June 2026 💫
meetings.cshl.edu/meetings.asp...
Bringing together experiments, computation and theory to explore dynamic, multi-scale cellular organization and function!
1/3
10.10.2025 09:23
👍 24
🔁 7
💬 2
📌 0

Nature Biomedical Engineering is hiring again! This time we're hoping to add an editor with machine learning expertise to the team (although we are open to those with other relevant expertise!) Deadline is Oct 20. Please RT! springernature.wd3.myworkdayjobs.com/SpringerNatu...
24.09.2025 18:36
👍 17
🔁 18
💬 1
📌 2

📢We're #hiring Group Leaders!
Apply to lead a lab at Janelia & advance biology using theory, computational modeling & machine learning.
🔹5-year renewable appointment
🔹Pioneer new tools & approaches
🔹Collaborate across disciplines
Apply by Nov. 4👉 https://janelia.link/groupleader
26.08.2025 19:00
👍 63
🔁 65
💬 0
📌 2
We are searching for an enthusiastic scientist to join our team and establish annual killifish as a model system for early developmental biology and biophysics! @embl.org @embldbunit.bsky.social
Apply here:
embl.wd103.myworkdayjobs.com/EMBL/job/Hei...
Please re-post 🙏
08.08.2025 09:01
👍 38
🔁 26
💬 2
📌 2

BIG start to #mmc2025 with Assaf Zaritsky giving an awesome opening plenary.
@royalmicrosoc.bsky.social @eurobioimaging.bsky.social @globias.bsky.social @bioimaginguk.bsky.social @globalbioimaging.bsky.social
01.07.2025 08:22
👍 19
🔁 4
💬 1
📌 2
Sorry, I meant @leeat-keren.bsky.social and Yael Amitay....
09.05.2025 09:15
👍 0
🔁 0
💬 0
📌 0

A creative and technically challenging idea led by @zamir_amos, with key contributions from Yuval Tamir and @yaelam75. This would not have been possible without the great collaboration with @leeat_keren! And thanks to @WellcomeLeap ΔTissue for funding!
16/n
09.05.2025 09:02
👍 0
🔁 0
💬 1
📌 0

Our results suggest that the local organization of a few cells in discriminative motifs are emergent properties that may define an intermediate spatial scale driving tissue function
14/n
09.05.2025 09:02
👍 0
🔁 0
💬 1
📌 0

CISM’s unsupervised motif enumeration followed by supervised context-dependent selection distils the discriminative motifs from the huge and noisy landscape of all putative sub-networks of intercellular interactions and provides discrimination along with interpretability
13/n
09.05.2025 09:02
👍 0
🔁 0
💬 1
📌 0
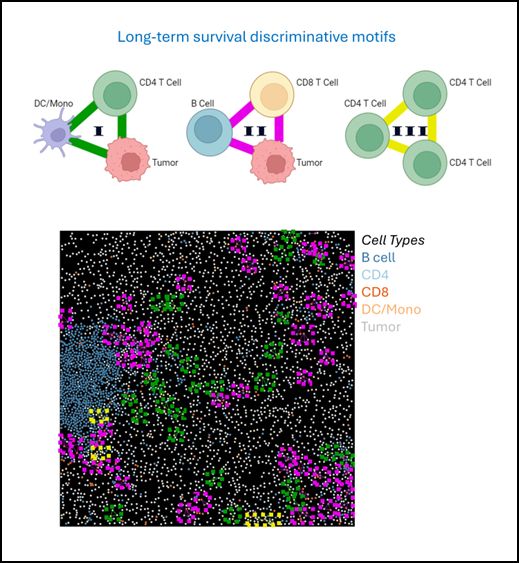
To demonstrate the general applicability of CISM for identifying local cellular structures linked to disease states, we applied it to investigate the tumor microenvironment in a human cohort of Breast Cancer (TNBC) patients, comparing short-term and long-term survivors.
12/n
09.05.2025 09:02
👍 0
🔁 0
💬 1
📌 0

The same motifs can be differentially analyzed according to different context-dependent selection of the discriminative motifs. We demonstrate this by investigating the immune microenvironment of NP versus PN (metastatic lymph nodes that did not develop distant metastases)
11/n
09.05.2025 09:02
👍 0
🔁 0
💬 1
📌 0

Spatial interpretation of motifs localization patterns reveals an association between (local) motifs to (global) tissue compartments highlighting the potential contribution of CISM to multi-scale analysis and interpretation
10/n
09.05.2025 09:02
👍 0
🔁 0
💬 1
📌 0

The spatial arrangement of cell types within the discriminative motifs contributed to disease state classification, indicating that the specific intra-motif cell-cell interactions are sensitive markers for disease state beyond their cell type composition
9/n
09.05.2025 09:02
👍 0
🔁 0
💬 1
📌 0

Exploring the landscape of the discriminative motifs-induced cell distribution and motifs-induced pairwise cell-cell interactions revealed differential composition of cell type and cell-cell interactions
8/n
09.05.2025 09:02
👍 0
🔁 0
💬 1
📌 0



















