Now published: our work using phylodynamics from surveillance data to quantify and experimentally validate the fitness impact of antibiotic resistance determinants & how this changes with patterns of antibiotic use: www.nature.com/articles/s41...
Now published: our work using phylodynamics from surveillance data to quantify and experimentally validate the fitness impact of antibiotic resistance determinants & how this changes with patterns of antibiotic use: www.nature.com/articles/s41...
How does treatment induced antibiotic resistance happen in real-world infections? We analysed 25k Pseudomonas isolates from 180 patients in a clinical trial to find out! TLDR: The ecological and evolutionary paths are surprisingly diverse & complex even in patients receiving identical treatment…

Great work again from @alexmsalmeida.bsky.social et al highlights uncultured genus CAG-170 (Oscillospiraceae) to be central node in healthy co-abundance networks in >11k gut metagenomes, with predicted B12 biosynthesis/cross-feeding functions possibly involved.
www.sciencedirect.com/science/arti...

🥁 NEW
Our study on infant gut microbiome and #straintransmission is now published in @nature.com:
doi.org/10.1038/s415...
1/10

Our team wrote a review for Gut Microbes on the role of the gut #microbiome in modulating enteric infections. Great team effort by Qi Yin, @samriddhigupta.bsky.social and Efrat Muller: www.tandfonline.com/doi/full/10....

Grateful to share our paper on gene-specific selective sweeps in human gut microbiomes, now out in Nature! It has been a joy to work with @rwolff.bsky.social, whose insights and hard work made this possible.
www.nature.com/articles/s41...

How do interactions with resident nasal microbiota shape colonisation resistance to MRSA?
Excited to share this preprint in collaboration with @mboum.bsky.social @anaellefait.bsky.social Silvio Brugger and Alex Hall.
www.biorxiv.org/content/10.6...

Now available online: the new 2.0 version of gutSMASH, with capabilities to detect 12 new types of catabolic gene clusters relevant to gut microbiome ecology, as well as predictions of their regulation through transcription factor binding site detection. www.sciencedirect.com/science/arti...

New PhD project, supervision by @alanmcn1.bsky.social and me.
'Nationwide Clostridioides difficile population dynamics'
www.findaphd.com/phds/project...
Deadline for applications 9 Jan 2026, UK students only.

Thrilled to share that our new paper is out now in @natcomms.nature.com 🎉
Huge congrats to @vhrcabral.bsky.social and @karinaxavierlab.bsky.social on this major work.
We found that our Klebsiella ARO112 can break the antibiotic/inflammation cycle in an IBD model

Diagram illustrating "feed the enemy's enemy": ampicillin inhibits Pseudomonas aeruginosa. Citrobacter freundii acidifies the environment, further inhibiting P. aeruginosa.
New preprint from our lab www.biorxiv.org/content/10.1...! Andrea Dos Santos and Clément Vulin combine experiments and models showing how adding glucose can strengthen negative interactions between microbial species. This can be used in tandem with antibiotic treatment to inhibit pathogens!

Long read Metagenomics, #phage and #prophage in the gut by Ami Bhatt's group. Beautiful data showing changes in phages over two years
#phagesky
www.nature.com/articles/s41...

🚨Preprint alert - this is a big one! We transfer the revolutionary power of TnSeq to bacteriophages.
Our HIDEN-SEQ links the "dark matter" genes of your favorite phage to any selectable phenotype, guiding the path from fun observations to molecular mechanisms.
A thread 1/8

New PhD position in my lab at @uniofbath.bsky.social (with both @tweethinking.bsky.social & Dr Bethan Littleford-Colquhoun)!
We're looking for someone keen on bioinformatics and microbiome evolution.
Important info below on eligibility & URSA competition funding👇
www.findaphd.com/phds/project...
Super excited that the bulk of my PhD work is now preprinted! Here we used whole-community competition, or coalescence, experiments to quantify selection acting on genetically diverged strains within larger communities. (1/n)
www.biorxiv.org/content/10.1...

Our paper “Global dissemination of npmA mediated pan‑aminoglycoside resistance via a mobile element in Gram‑positive bacteria” is now in @natcomms.nature.com. Part of my freshly defended PhD, so doubly happy! 😄🎉
🧵 (1/14)
www.doi.org/10.1038/s414...

Can we leverage bacterial competition for targeted replacement of harmful strains? Maybe! Our recent piece in @natmicrobiol.nature.com provides a theoretical framework and a set of experiments to show what it might take: www.nature.com/articles/s41...
New paper out 🎆 When an antibiotic-resistant E. coli strain lands in our gut microbiome, whether it will get established or not depends on the ecological context. We studied how other microbes, nutrients, and antibiotic exposure shape its fate.👇

So happy to share this! Bacteriocins were first discovered over 100 years ago, but what do they actually do? We look at >1000 bacteriocin plasmids and find links to virulence and antimicrobial resistance, and frequent bacteriocin sharing in Enterobacteriaceae.
www.nature.com/articles/s41...

🐝🦠 New paper: rdcu.be/eOf7A
Phages may drive microbial diversity, yet we often don’t even know how phages & bacteria correlate in nature. Our new study tackles this in the honeybee gut, thanks to the great work of PhD student @malickndiaye.bsky.social at @dmf-unil.bsky.social @fbm-unil.bsky.social

New(ish!) paper on how within-host competition and antibiotic resistance shape the fitness of Streptococcus pneumoniae serotypes, out in August in Plos Biology. journals.plos.org/plosbiology/...
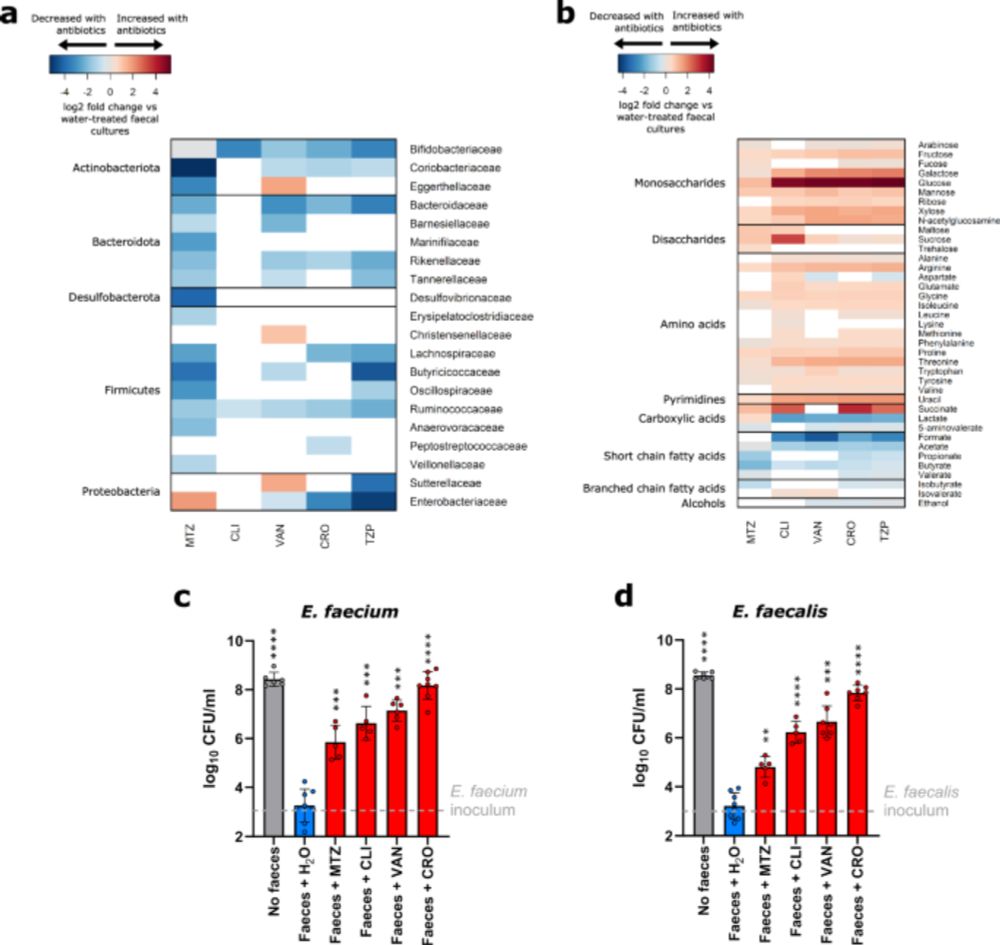
So excited to share a new paper from my lab just published in Nature Communications! We showed that vancomycin-resistant enterococci (VRE) occupied distinct intestinal niches in the antibiotic-treated intestine. doi.org/10.1038/s414... Amazing work by first author Olivia King and colleagues!

OUT NOW - Long-read metagenomics method for strain tracking after FMT #microsky www.nature.com/articles/s41...

Deciphering microbial spatial organization: insights from synthetic and engineered communities url: academic.oup.com/ismecommun/a...
Our high-precision metagenomic strain caller, PHLAME, is now published in Cell Reports!! www.cell.com/cell-reports...
PHLAME works on tough sample types -- including those with coexisting strains of a species and low depth.

Mechanisms of microbiome assembly and approaches to uncover the ecological forces driving it.
Deriving ecological models solely from observational data limits our ability to understand mechanisms driving microbiome assembly. This #mSystems articles explores how experimental approaches can address these challenges. asm.social/2wj
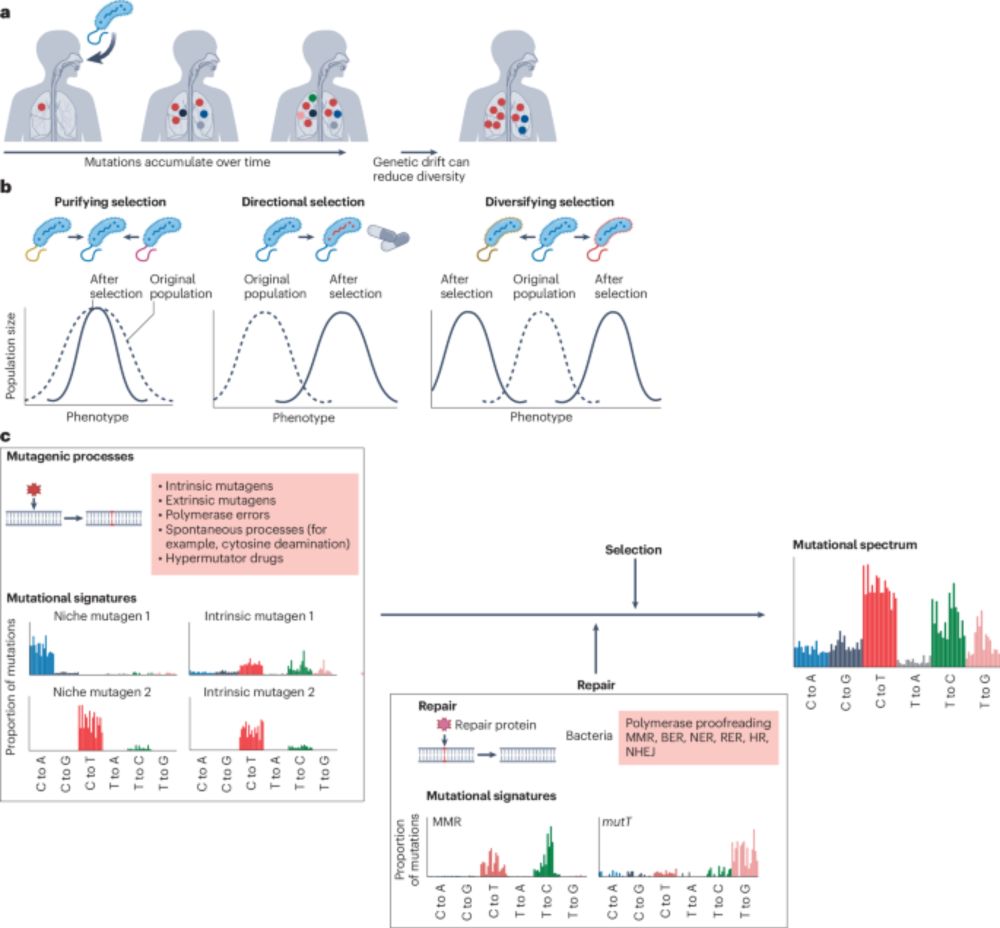
#Review
A discussion on the processes driving bacterial evolution and emergence of pathogenesis within hosts, the importance of understanding within-host genetic diversity, and the implications for transmission analysis and infectious disease control.
#MicroSky 🦠
www.nature.com/articles/s41...
Our paper demonstrating that within-species warfare interactions are ecologically important on human skin is now published in Nature Micro! www.nature.com/articles/s41...
This is such a cool read on the fascinating topic of nutrient competition in microbiomes😍 I couldn t agree more with «A key challenge in understanding microbiomes is that the species composition often differs among individuals, which can thwart generalization. »
Our work on the facial skin microbiome of non-human primates is out in mSystems!
We show there is no close relative of Cutibacterium on the faces of gorillas and chimps at the Lincoln Park Zoo, furthering the mysterious origin of the dominant human skin colonizer.
journals.asm.org/doi/10.1128/...