@epfl-brainmind.bsky.social @epfl-ai-center.bsky.social @scadsai.bsky.social
@epfl-brainmind.bsky.social @epfl-ai-center.bsky.social @scadsai.bsky.social
Very happy that MuViT, our (CVPR 2026!) work on learning across spatial scales in microscopy w transformers, is out! Check Martin's thread for a nice walkthrough :)
Big thanks to my advisors @maweigert.bsky.social @gioelelamanno.bsky.social!
🖥️: github.com/weigertlab/muvit
📜: arxiv.org/abs/2602.24222
Excited to share our new paper (CVPR 2026 🚀): "MuViT: Multi-Resolution Vision Transformers for Learning Across Scales in Microscopy" which enables local predictions to use global context.
Great work led by @albertdm.bsky.social & another fun collab w @gioelelamanno.bsky.social! @scadsai.bsky.social

📣We’re back 🔬👀🖥️!
#CBIAS2026 returns 23–24 November 2026:
Join the bioimage analysis community to discuss advances in quantitative imaging & computational methods for image analysis. Great opportunity for early-career analysts & microscopists to present and connect.
#CBIAS2026 #BioimageAnalysis
Last call! This is a great opportunity for those interested in learning and applying SOTA ML/DL models to microscopy data with the leading experts in the field.
Personally super excited to be a lead TA along with the amazing @afoix.bsky.social and @edhirata.bsky.social!
Hope to see you there :)

Now speaking #CBIAS2025 @albertdm.bsky.social from @maweigert.bsky.social introducing spotiflow www.nature.com/articles/s41...
Had a blast at #CBIAS2025! Great science (like the incredible ShapeEmbed talk from @afoix.bsky.social), amazing people and obviously tons of beautiful images :) Already looking forward to coming back next year!

We’ve upgraded ShapeEmbed 🎉 ShapeEmbedLite decodes latent codes via an MLP to guarantee valid EDMs, making it lighter and ideal for small microscopy datasets or limited compute. Hear more at my BIC workshop talk or poster at #ICCV2025! Try it out at github.com/uhlmanngroup...

🧠 The Lipid #Brain Atlas is out now! If you think #lipids are boring and membranes are all the same, prepare to be surprised. Led by @lucafusarbassini.bsky.social with Giovanni D'Angelo's lab, we mapped membrane lipids in the mouse brain at high resolution.
www.biorxiv.org/cgi/content/...
You may have heard me talk about learned shape representations for a long time, but it took @afoix.bsky.social's hard work to bring it to life 🍾 Catch her at @neuripsconf.bsky.social to find out more!!

Happy to share that ShapeEmbed has been accepted at @neuripsconf.bsky.social 🎉 SE is self-supervised framework to encode 2D contours from microscopy & natural images into a latent representation invariant to translation, scaling, rotation, reflection & point indexing
📄 arxiv.org/pdf/2507.01009 (1/N)
(1/14) I’m happy and proud to introduce: SpinePy – a framework to detect the "spine" of gastruloids and measure biological and physical signals in a local dynamic 3D coordinate system. www.biorxiv.org/content/10.1...

Happy to share our work on uMAIA, a framework for building metabolomic atlases from mass spectrometry imaging.
With uMAIA, we mapped the lipidome of zebrafish development, uncovering spatially organized metabolic programs .
www.nature.com/articles/s41...
#developmentalbiology #MSI #lipidtime

A new extension! Rémy made a wrapper of @albertdm.bsky.social + @maweigert.bsky.social 's Spotiflow (2/3)
forum.image.sc/t/qupath-ext...
Cell tracking is never perfect, and it's important to understand the types of errors your solution contains.
Here is my stab at this: Divisualisations in @napari.org. github.com/bentaculum/d...
You spin tracks out upwards from the playing video. Green edges are correct, FP in magenta, FN in cyan.
Working in Martin’s group is an amazing experience, don’t hesitate to apply and reach out if you have any questions!

Out today in @natmethods.nature.com : Spotiflow, our transcript localization method for imaging-based spatial transcriptomics. Led by amazing PhD student @albertdm.bsky.social, joint work w @gioelelamanno.bsky.social at EPFL / @scadsai.bsky.social
www.nature.com/articles/s41...
rdcu.be/epIB7
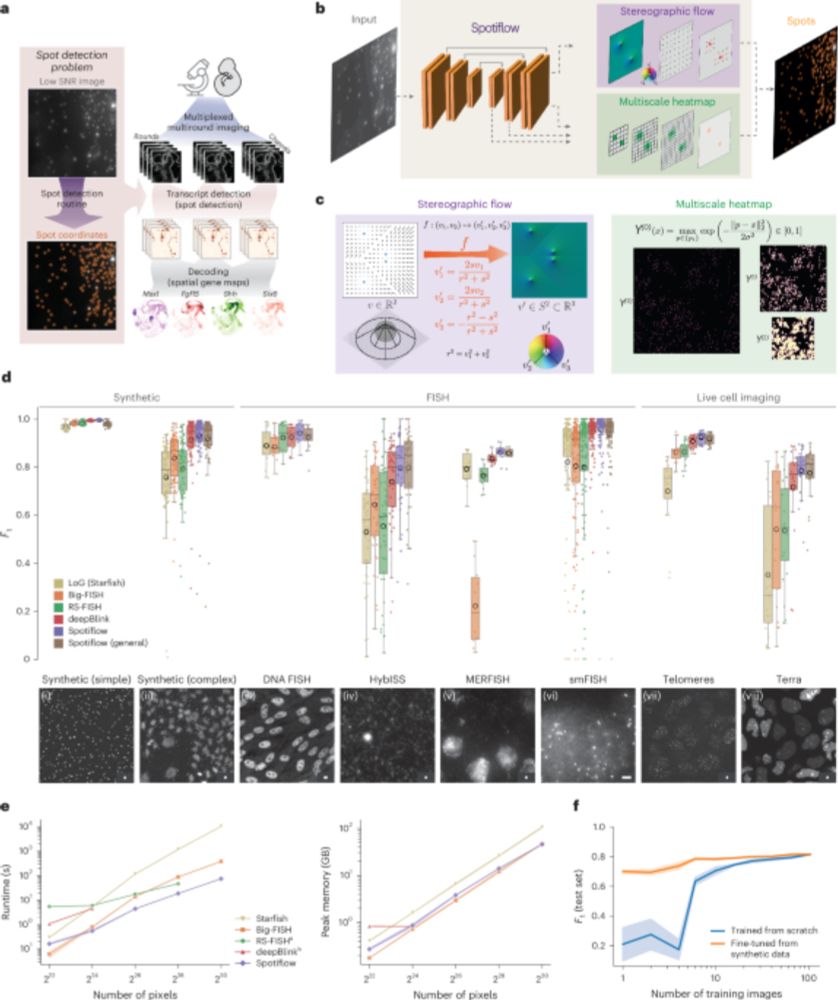
A nice advance for imaging-based spatially resolved transcriptomics from the Weigert and La Manno labs. Spotiflow uses deep learning for subpixel-accurate spot detection in diverse 2D and 3D images. www.nature.com/articles/s41...
A massive thanks to all the incredible people who contributed to this work, it's been an amazing ride! And a shoutout to all the people who have tried already tried Spotiflow and provided feedback, it's truly great to see the community using it :)
(N/N)

For more details on using Spotiflow, check our repository at github.com/weigertlab/s...! We are very excited to see what uses cases people will find for Spotiflow!
(9/N)

Results showing the effect of the number of training images on model performance for training from scratch and fine-tuning
We also offer several out-of-the-box pre-trained 2D/3D models. Fine-tuning a model on your own data is easy and accessible if you're not satisfied with the pre-trained ones, and we show Spotiflow is not particularly data hungry, so no need to spend a lot of time annotating!
(8/N)
We developed Spotiflow with the mindset of making it useful and usable, and as such it comes in different flavors: one can use it as a Python API, as a CLI or with a GUI using our @napari.org plugin! Hopefully more integrations will come soon ;)
(7/N)

Overview of EASI-FISH data + pairs of insets containing raw (left) and Spotiflow detections (right)
To be ready to handle today’s massive spatial biology data, we built a multi-GPU setup that e.g. processes a cloud-stored 159 GB EASI-FISH volume in under 10 minutes on 8×A100 GPUs. This can easily be deployed seamlessly on any modern compute cluster!
(6/N)
In a single molecule tracking experiment w/ Eftychia Kyriacou and Joachim Lingner from EPFL, Spotiflow allowed us to improve results by changing the input detections to Trackmate, showing that Spotiflow can be used as a drop-in replacement in existing workflows!
(5/N)
The context-awareness of Spotiflow allows for precise detection in challenging SNR conditions. This includes avoiding artifacts due to e.g. autofluorescence as we show with the beautiful P. dumerilii volumetric smFISH data from the Arendt and @ilastik-team.bsky.social teams @embl.org
(4/N)

Benchmarking results for different spot detection methods
Spotiflow achieves SOTA results on several datasets (iST, live imaging, DNA FISH) while being faster than commonly-used spot detection methods when used out-of-the-box on large-scale data, both in 2D and 3D!
(3/N)
Given an image or volume, Spotiflow uses a neural network to predict multiscale Gaussian heatmaps and the “stereographic flow” — a smooth embedding that maps the nD nearest-spot vector field to the (n+1)D sphere, avoiding issues at infinity and enabling precise localization
(2/N)
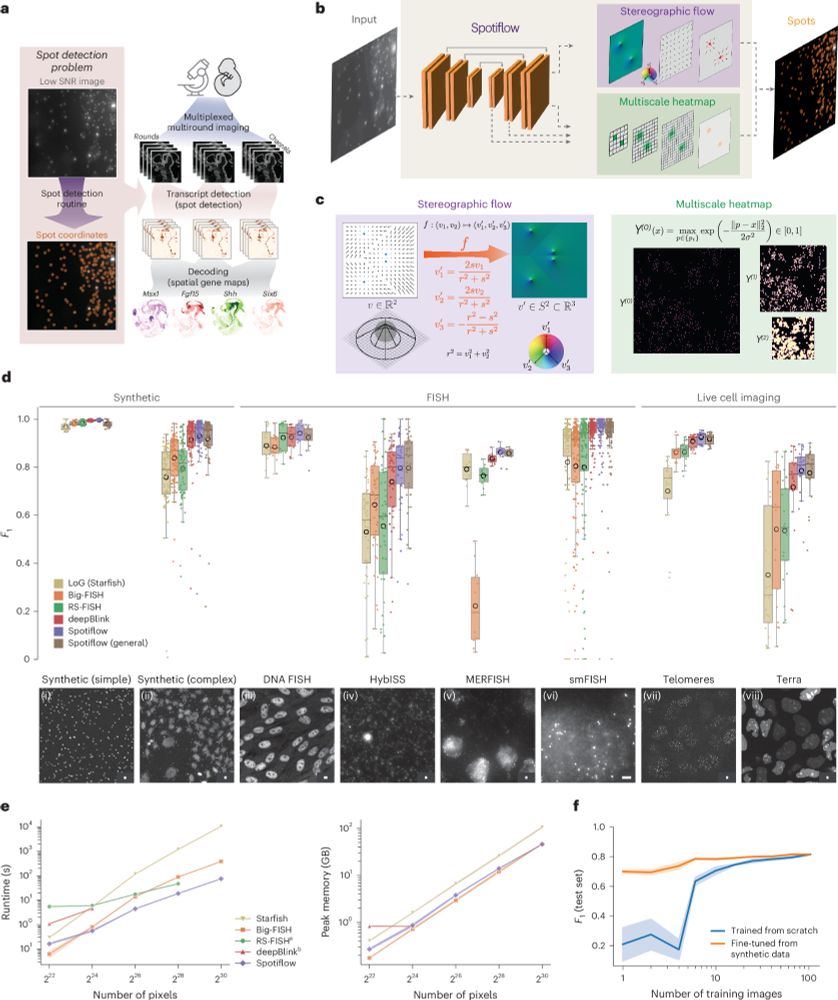
Spotiflow, our deep learning based spot detection method for microscopy, is now published in @natmethods.nature.com!
Since the pre-print, we have added many features, notably native 3D detection!
@maweigert.bsky.social @gioelelamanno.bsky.social @epfl-brainmind.bsky.social
Paper: rdcu.be/epIB7
(1/N)

Get to know Spotiflow, an AI method that improves large-scale transcript localization by combining deep learning with a novel geometric detection approach.
Learn more:
👉 scads.ai/transcript-d...
Find the publication in Nature Methods:
👉 www.nature.com/articles/s41...

How can AI help you to analyze your microscopy images?
Find out at the "Machine Learning for Microscopy Image Analysis" course at the @mblscience.bsky.social in Woods Hole from Aug 22-Sep 6!
Applications are due 🚨 April 30 🚨
Read on for more! 👇
www.mbl.edu/education/ad...