Reposting! We’re going to do some amazing science and have a lot of fun doing it!
Reposting! We’re going to do some amazing science and have a lot of fun doing it!

Come do a PhD with me and @trevor-lithgow.bsky.social at @monashuniversity.bsky.social ! We’re at the leading edge of pathogen biology using cutting edge Cryo-EM and AI-driven genetic screens to uncover the mysteries of outer membrane biology. Reach out to me for a chat!
macsys.org/phd-scholars...

This year Hartland Oration was delivered by Thomas McLean from @wehi-research.bsky.social, a great talk on how does the structure of DNA sliding clamp enable gene silencing on a multidrug-resistance plasmid. 👏 #lorneiandi
First results chapter from Kelly-Rose Tulley’s PhD is published. It was a great team effort to get this finished. This work supports previous results on S. coelicolor OrrA and suggests its function is highly conserved. #microsky #streptomyces 1/2
Too excited by the science to include the link to the paper! Here it is: journals.asm.org/doi/10.1128/...
After all that, i just want to thank all the amazing co-authors but particularly @ainsley-beaton.bsky.social for helping me get to the bottom of this story, and my PhD supervisor @matthutchings.bsky.social for always believing in the project!

This leads to the activation of HtrA-family dual function chaperone/proteases that can act to restore the cell to homeostasis! To our knowledge, this is the first proposal of a sensing model for a bacterial secretion or envelope stress response system! (6/7)
Finally, through ChIP-seq, TMT-proteomics and qRT-PCR we managed to link it all. Together with CssRS, CutRS detect secretion stress through the formation (or lack thereof) of the conserved disulfide bond in their sensor kinase sensor domain. (5/n)

Finally, with the help of the amazing @ainsley-beaton.bsky.social, we had a breakthrough. Using a combination of modelling and phylogenetics we discovered that the CutS sensor kinase had a highly conserved dual cysteine motif in its extracellular sensor domain, suddenly we had some clues! (4/n)

My favourite piece of data in the whole paper came after we quantified the biomass difference between the WT and ∆cutRS mutant on glucose media, showing it is around 4x more massive. It also uses up all the glucose in the plate, close to gram, in just 9 days! (3/n)

This mutant also overproduced the antibiotic chloramphenicol, leading to a very potent anti-Gram-postitive activity on plate assays. However, this still didn't give us any clues as to what linked CutRS to the phenotype. (2/n)

Delighted to see the main work from my PhD finally published in @mbio.bsky.social! It all started with the observation that deleting the cutRS two-component system in S. venezuelae caused this amazing explorer phenotype in the presence of glucose. But what was going on?! (1/n)

📄 Delighted to share that the preprint of my main PhD work is out now!
👏 A huge thank you to everyone involved, @tommclean.bsky.social, Govind Chandra, Ngat Tran (both not on BlueSky) and especially my PhD supervisor @tunglejic.bsky.social 👏
🧵 below!
www.biorxiv.org/content/10.6...
Check out this work by the amazing @leahmcphillips.bsky.social from during her PhD!!

Our preprint is now published in PNAS! This came together thanks to a great collaboration with Antoine Hocher and a strong team effort from the Le Lab. Thank you to the reviewers and to everyone who helped improve it. I hope ParB aficionados will enjoy it.
www.pnas.org/doi/10.1073/...
Wooo congrats!!

#straya
Bin Chicken: targeted metagenomic coassembly for the efficient recovery of novel genomes.
www.nature.com/articles/s41...
When people celebrate the individual genius of folks in science, they should also
mourn the collective loss of genius of folks who were actively discouraged or disadvantaged from a career in science because of the same person(s)

new preprint from our group & Antoine Hocher: www.biorxiv.org/content/10.1...
A fantastic collaboration with Antoine, with Jovana Kaljevic' initiated the collaboration and drives the project.
Versatile NTP recognition and domain fusions expand the functional repertoire of the ParB-CTPase fold beyond chromosome segregation https://www.biorxiv.org/content/10.1101/2025.10.10.680097v1
Yes Miguel!!! Huge congrats that’s amazing news! 🎉🎉🎉
Ever wondered why some antibiotics are made by Streptomyces on agar plates but not in liquid cultures? Read this work on redox control of antibiotic biosynthesis. Led by katienoble241.bsky.social and Rebecca Devine, and in collaboration with @barriewilks.bsky.social
journals.asm.org/doi/10.1128/...

After 8 wonderful years in Norwich, 3 at UEA and 5 at the JIC it’s time to say goodbye. I will miss each and every amazing person I’ve had the pleasure of working with or meeting here. It truly is a fine city.

Nice to see our recent work on the fascinating KorAB system be featured on the back of @johninnescentre.bsky.social Advances!
Read the full paper here: www.nature.com/articles/s41...
Check out our lab's new work on a CTP-independent ParABS system. Some fantastic work from Kirill et al!
Have a read of this simply phenomenal work on the weird and wonderful GTAs from star Fellow (and office mate) Emma Banks et al
Fantastic work Trevor!
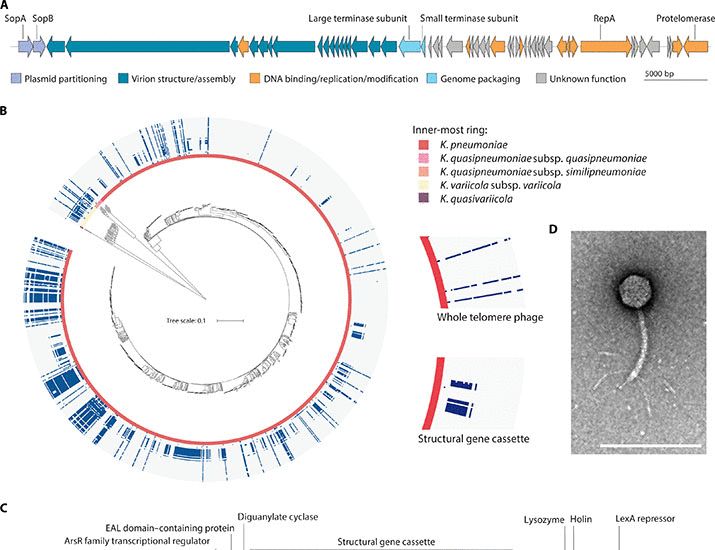
"That telomere phages are so prevalent means that they are a selective force, one that we know little about. We now want to understand how the telomere-toxin is secreted and also understand how this ‘telocin’ wheedles its way into unsuspecting bacterial neighbors”
www.science.org/doi/10.1126/...
The paper is now live: academic.oup.com/mbe/article/...
Absolutely delighted to play a role in this fascinating story!
The second great paper concerns the discovery in Eva Top's group that an enigmatic protein encoded by IncP-1beta plasmids modulates the IncP-1 circuitry in E. coli to make the plasmid a burden in this host: Elg et al recently accepted in Molecular Biology and Evolution doi.org/10.1093/molb...