📢 Job alert (3/3)
🌱 The goal is to build a team with diverse backgrounds, so please don’t be discouraged if you don’t tick all the boxes!
www.leibniz-lsb.de/fileadmin/do...
www.leibniz-lsb.de/fileadmin/do...
📢 Job alert (3/3)
🌱 The goal is to build a team with diverse backgrounds, so please don’t be discouraged if you don’t tick all the boxes!
www.leibniz-lsb.de/fileadmin/do...
www.leibniz-lsb.de/fileadmin/do...
📢 Job alert (2/3)
💼🧬 I’m currently looking for two motivated PhD students to start around May. If you or anyone you know is excited about graphs, (probabilistic) ML, omics data, and applications to food biology, please apply/reach out and spread the word.
📢 Job alert (1/3)
In April, I will be starting my research group at the Leibniz Institute for Food Systems Biology at TUM.
💻🍣 Our work will focus on developing computational methods to integrate and analyze omics data in a food biology context

We’re excited to be back at the Heidelberg spatial omics seminar series today 😍
I think that is the core of the peer-review crisis. If we were to only do peer review of the scientific work that is both important enough and non-trivial, we would all save a lot of time. It will obviously be a challenge to decide what needs to be reviewed, I acknowledge that.

View from the hotel room

Poster session 2024, with Valentina Boeva, Constantin Ahlmann-Eltze and others

Wednesday afternoon hike incl. swim in the mountain river

Another view from the hotel room
Apply for the Ascona workshop "Statistical and AI methods for multi-modal multi-scale modeling of biological systems", 28 Jun-3 Jul 2026 on Monte Verità, Lago Maggiore at the foot of the Swiss Alps.
ascona2026.sciencesconf.org

“What a century, uh?” “Captain, it’s just one quarter”

A new lab preprint is up: Work led by our MSc student Jan Sprengel comparing Interventional Causal Structure Learning Algorithms for Gene Regulatory Network Inference from single cell CRISPR perturbation screens 👇 www.biorxiv.org/content/10.6...
We have been working with Michal Klein on pushing a module to train *flow matching* models using JAX. This is shipped as part of our new release of the OTT-JAX toolbox (github.com/ott-jax/ott)
The tutorial to do so is here: ott-jax.readthedocs.io/tutorials/ne...

Look at the numbers! They are great! Good numbers, bold numbers, we have the best numbers!
It's amazingly fast (< 𝟏𝐡 𝐟𝐨𝐫 𝐭𝐡𝐞 𝐟𝐮𝐥𝐥 𝐩𝐢𝐩𝐞𝐥𝐢𝐧𝐞, 𝐭𝐫𝐚𝐢𝐧𝐞𝐝 𝐨𝐧 𝐚 𝐠𝐚𝐦𝐢𝐧𝐠 𝐥𝐚𝐩𝐭𝐨𝐩) and high-quality.
Another Topological Deep Learning success story, coming soon to #NeurIPS2025!
🖥️ github.com/aidos-lab/in...
📜 arxiv.org/pdf/2410.18987
4/n
Extremely honoured to receive the Bayer foundation early excellence in science award. Thanks to all my team, colleagues, collaborators and mentors for making this possible!
My take: Engineering *is* important. Ideas are cheap. Novelty is overrated. "Hyperparameter tuning" is a cynically dismissive term for something that is in fact the real, difficult problem.

Nikolai Koehler goes over the materials of the first part of the course.

Panoramic view of the EMBO course classrom while Nikolai Koehler is talking.

Wangjun Hu goes throught the materials of the second part of Practical session 5.
After the coffee break ☕, we continue with the second part of practical session 5, with Nikolai Köhler and Wangjun Hu 🎓 going through the different materials: from storing spatial-omics data 💽 to learning spatial patterns using #MEFISTO 😎



Welcome to day 4 of #EMBOBioImage 🖥️
👉Happening right now: practical session 5, spatial statistics and omics integration, led by @brittavelten.bsky.social, assisted by Anna Klemm.
I always find it funny that we obsess over sample size, yet peer review relies on the arbitrary opinions of just 2/3 people. That would be fine if editors were truly engaged, but given the workload, they don’t have the time, or don’t care enough. Anyway, I’m just glad it finally got published! 👏
@embl.org PhD programme just open - many interesting projects, including two with us @saezlab.bsky.social @ebi.embl.org - one on #host-microbiome interactions using #spatial omics, and one on #AI in #proteomics 👇

Very happy to share our latest publication unravelling how the #ERstress sensor ATF6 stirs up trouble (and tumors) in #colorectalcancer. ATF6 drives CRC by reshaping lipid #metabolism and the intestinal #microbiota. A huge thank you to all co-authors! #lifesciences
www.nature.com/articles/s42...

We spent a year writing this review of low-dim embeddings and arguing about things like epistemic roles and best practices :-) 20+ authors are all participants of the Dagstuhl seminar we held last year: www.dagstuhl.de/24122. Led by @alexandr.bsky.social and Cyril de Bodt.
arxiv.org/abs/2508.15929
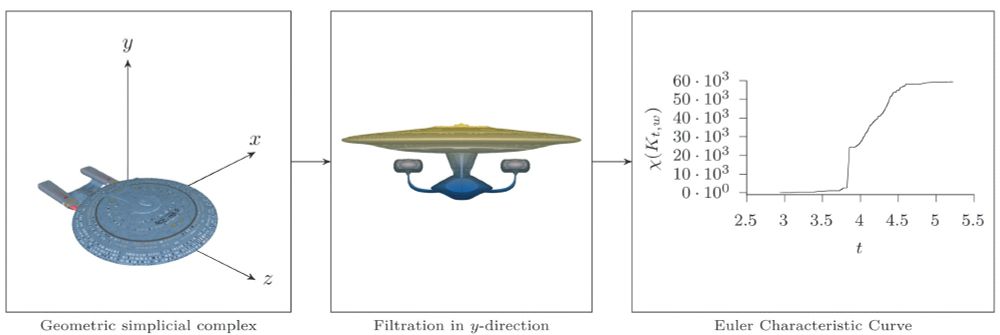
An overview of the Euler Characteristic Transform, illustrated for a single Euler Characteristic Curve, i.e., the calculation of Euler characteristics along a single direction.
#MachineLearning loves data. #Topology loves shape.
Let's bring them together!
👉The Euler Characteristic Transform (ECT) bridges both worlds, as I outline in a recent article in the Notices of the
@amermathsoc.bsky.social
Engage!
🧵1/n


Exciting kick-off meeting for our spatial omics seminar in and around Heidelberg organised with @tanevski.bsky.social and @denisschapiro.bsky.social. Was fun to see all the cool research happening here and connect. Reach out to us if you missed out and would like to join the next meetings!

🚀 Job opportunity: Bioinformatics Software Engineer
Join a young and passionate team at Heidelberg University to advance bioinformatics software that drive biological discovery.
Learn more & apply 👉 shorturl.at/GCpg7



We're excited to be at the Kick Off Heidelberg Spatial-Omics Lunch Seminar Series with our very own @chiaraschiller.bsky.social presenting some of her PhD work!

🌟We're glad to announce that the brand new EMBL Conference #EMBLSpatialBiology is now open for registration ➡️ https://s.embl.org/spb25-01-bl
Submit your abstract by 8 July and be part of the rapidly growing spatial biology community!
🗓️ 14 – 17 October
📍 EMBL Heidelberg and Virtual
#ESSB2025
JOB OPPORTUNITY: Join us as a scientific programmer to advance tools for multi-omics data at Heidelberg University. Details can be found at shorturl.at/OcnOa and please spread the word!