
New article from me and collaborators just dropped.
hbr.org/2026/03/peer...

New article from me and collaborators just dropped.
hbr.org/2026/03/peer...
Five years ago, we released FLIP. The core question was: can ML models for protein fitness prediction generalize in the ways that actually matter for protein engineering, i.e. low data, extrapolation to more mutations, out-of-distribution sequences?

We made FLIP2, a protein fitness benchmark spanning seven new datasets, including enzymes, protein-protein interactions, and light-sensitive proteins, as well as splits that measure generalization relevant to real-world protein engineering campaigns.

FoldMason is out now in @science.org. It generates accurate multiple structure alignments for thousands of protein structures in seconds. Great work by Cameron L. M. Gilchrist and @milot.bsky.social.
📄 www.science.org/doi/10.1126/...
🌐 search.foldseek.com/foldmason
💾 github.com/steineggerla...

For no reason, I remembered today that I too once got to take a picture holding Nobel Prize that I didn't earn
Never believe the fuckers who say "why bother, they already won"
Updated preprint on the roadmap to predicting macromolecular ensembles with @bonomimax.bsky.social! Four key issues:
1) Structural ensembles are defined inconsistently across disciplines.
2) No single experimental technique alone can fully capture structural ensembles
arxiv.org/abs/2505.01919
In Boston, every day is No Kings Day
😍 Boston, you are beautiful. #NoKings

My linocut portrait of astronaut Mae Jemison in her flight suit, holding her helmet, with the globe of Earth above and to left.
Happy birthday to American astronaut Mae Carol Jemison (born October 17, 1956), a physician who became the first Black woman to travel in space when she went into orbit aboard the Space Shuttle Endeavour for NASA, on September 12, 1992! 🐡🧪🧑🏾🔬 #histsci She also has a B.S. in chemical engineering, 🧵1/n
Want to join the BioEmu team at MSR AI for Science? We have multiple intern (jobs.careers.microsoft.com/global/en/jo...) and one fixed-term postdoc (jobs.careers.microsoft.com/global/en/jo... positions available! Please share with potential candidates :)

Figure 3 from the paper with the caption: "Role of machine learning in de novo design of IDRs. (A) Machine-learning models can be trained on diverse data sources, from molecular dynamics simulations to annotations of cellular localization and protein structures from the Protein Data Bank. (B) Often implemented as neural networks using sequence-encoded features as input, these models can initially be trained on a limited region of sequence space as surrogate models. Through active learning, additional simulations are performed during the design campaign to generate new data, and the surrogate model is retrained on the expanded dataset to progressively improve its accuracy. (C) Machine-learning models have been developed to predict biophysical observables, biological annotations, and protein structures. When combined, machine-learning models can be used to identify a set of sequences that strike a trade-off between multiple design objectives, defining a Pareto front."
New review on computational design of intrinsically disordered proteins 🖥️🍝 by @giuliotesei.bsky.social @fpesce.bsky.social & 👴
doi.org/10.48550/arX...

After years of complaining about cancel culture, the current administration has taken it to a new and dangerous level by routinely threatening regulatory action against media companies unless they muzzle or fire reporters and commentators it doesn’t like.

Tomorrow we're excited to host @sarahalamdari.bsky.social at Chalmers for the AI4Science seminar and hear about generative models for protein design! Talk at 3pm CEST. 🤩
For more info, including details on how to join virtually, please see psolsson.github.io/AI4ScienceSe...
@smnlssn.bsky.social


A compelling review of how ML/AI could help in the quest to find an enzyme for every reaction.
@jsunn-y.bsky.social @francescazfl.bsky.social Yueming Long @francesarnold.bsky.social
www.cell.com/cell-systems...
Join us this fall for the Chalmers AI4Science seminar series with this amazing line-up of speakers! All seminars held on-campus at Chalmers and streamed over Zoom.
See psolsson.github.io/AI4ScienceSe... for details!
Speakers:
@sarahalamdari.bsky.social
@micheleceriotti.bsky.social
& Prashant Singh

A United States Senate investigation has identified more than 500 credible reports of human rights abuses in US immigration detention since January, including alarming allegations of mistreatment of pregnant women and children.

AtomWorks is out! Building upon @biotite_python, we built a toolkit for all things biomolecules and trained RF3 with it. All open-source, test it via `pip install atomworks`!
AtomWorks: github.com/RosettaCommo...
RF3: github.com/RosettaCommo...
Paper: tinyurl.com/y2w4z65b
1/6

(1/7)
Training biomolecular foundation models shouldn't be so hard. And open-source structure prediction is important. So today we're releasing two software packages: AtomWorks and RosettaFold3 (RF3)
[https://www.biorxiv.org/content/10.1101/2025.08.14.670328v2](www.biorxiv.org/content/10.1...)


The @uwproteindesign.bsky.social's experimental pipeline behind models like RFdiffusion and ProteinMPNN:
- A rapid, scalable, pipeline for producing and characterizing proteins
- A demultiplexing protocol for converting oligopools to clonal constructs
Jason Qian @lfmilles.bsky.social Basile Wicky

Excited to share work with
Zhidian Zhang, @milot.bsky.social, @martinsteinegger.bsky.social, and @sokrypton.org
biorxiv.org/content/10.1...
TLDR: We introduce MSA Pairformer, a 111M parameter protein language model that challenges the scaling paradigm in self-supervised protein language modeling🧵
I don’t want the NIH to be fucking profitable, I want it to cure cancer and Alzheimer’s
We may have the chance to hire an outstanding researcher 3+ years post PhD to join Tarleton Gillespie, Mary Gray and me in Cambridge MA bringing critical sociotechnical perspectives to bear on new technologies.
jobs.careers.microsoft.com/global/en/jo...


In 1965, Margaret Dayhoff published the Atlas of Protein Sequence and Structure, which collated the 65 proteins whose amino acid sequences were then known.
Inspired by that Atlas, today we are releasing the Dayhoff Atlas of protein sequence data and protein language models.

Join us along the #BostonPride parade route to celebrate the resistance of our LGBTQIA+ community in the face of tyranny.
✊🏳️🌈🏳️⚧️🇺🇸
#NoKings
#50501movement
@50501movement.bsky.social
www.boston.com/news/local-n...
Massive crowd at Boston’s city hall plaza for the #HandsOff protest against Trump and Musk

A large paper clip with googly eyes glued to it. It’s standing up next to the track pad on a laptop due to magic aka magnetic forces
I accidentally discovered that there is a weird magnetic spot on my laptop that will make a paperclip stand up, so I had to add googly eyes and make my own personal Clippy
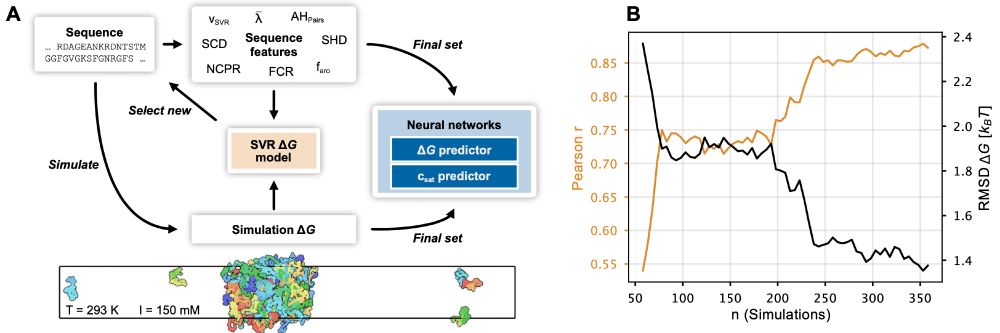
Our paper on prediction of phase-separation propensities of disordered proteins from sequence is now published:
www.pnas.org/doi/10.1073/...
The paper has been substantially updated compared to the preprint including new experimental data and using the neural network to finetune CALVADOS. 1/n

Protein dynamics was the first research to enchant me >10yrs ago, but I left in PhD bc I couldn't find big experimental data to evaluate models.
Today w @ginaelnesr.bsky.social, I'm thrilled to share the big dynamics data I've been dreaming of, and the mdl we trained w them: Dyna-1.
📝: rb.gy/de5axp