
Check out our new preprint in which we teamed up with the Walter and Chen labs to determine how transcription coordinates the folding and function of a tandem aptamer glycine riboswitch from B. subtilis: www.biorxiv.org/content/10.1...

Check out our new preprint in which we teamed up with the Walter and Chen labs to determine how transcription coordinates the folding and function of a tandem aptamer glycine riboswitch from B. subtilis: www.biorxiv.org/content/10.1...
Importantly, all the materials needed to perform this assay are commercially available, so the method is broadly accessible.
This has implications for the ability of pseudoknots to stabilize transcription pauses, which the Walter lab previously showed is possible (www.sciencedirect.com/science/arti...).
In addition to validating the method by assessing five RNA folding transitions in three RNA targets, we unexpectedly observed at least three pseudoknot base pairs forming within the RNA exit channel of RNAP.
This strategy is distinct from previous methods (Cotranscriptional SHAPE-Seq, SPET-Seq, TECprobe-VL), which measure cotranscriptionally folded RNA, but not cotranscriptional RNA folding - in our new approach, we directly assess how RNA structure changes as RNAP transcribes downstream.

In this study (www.nature.com/articles/s41...) we used a reversible transcription roadblocking strategy developed by the Lafontaine lab to sequentially position E. coli RNAP at two or three positions within a DNA template so that we can take fractions of the reaction for chemical probing.

Our paper describing a method for measuring cotranscriptional RNA folding using high-throughput RNA chemical probing is out @natcomms.nature.com!
Actually, I was off on the year of Opiate, so only Fear Inoculum counts (better album IMHO)
Tool - 10,000 Days and Fear Inoculum both qualify.

Here is our paper showing in vivo folding of nascent RNA right after synthesis, published now in @cp-molcell.bsky.social!
With @karlaneugebauer.bsky.social, @isaac-vock.bsky.social, @mattdsimon.bsky.social
www.cell.com/molecular-ce...
I'm excited to share our latest publication on co-transcriptional RNA folding in a eukaryote! Great work by Leo Schärfen with collaboration from Isaac Vock and Matt Simon! @leoschaerfen.bsky.social @mattdsimon.bsky.social
Take a peek here authors.elsevier.com/a/1kpxo_Oylr...
I'm excited to share our latest publication on co-transcriptional RNA folding in a eukaryote! Great work by Leo Schärfen with collaboration from Isaac Vock and Matt Simon! @leoschaerfen.bsky.social @mattdsimon.bsky.social
Here is the link that works! authors.elsevier.com/a/1kpxo_Oylr...
Yup. I use SABIAN cymbals and inquired about getting some prototypes through SABIAN's Sound Check program in January. I was told that the program was on hold for the US until the uncertainty regarding tariffs is resolved. Glad I'm already set up with all the catalog cymbals I want for now...

BOOM! US tariffs could crash the market for North American drummers, writes PIIE's Cullen Hendrix www.piie.com/blogs/realti...

Our paper describing our Transcription Elongation Complex Display platform for high-throughput cotranscriptional RNA functional assays is now online at @naturecomms.bsky.social: www.nature.com/articles/s41.... We used TECdisplay to assess the activity of >1M ZTP riboswitch variants!
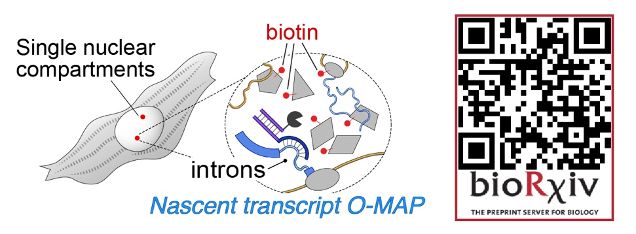
It's my great pleasure to present the next big preprint from SheqLab! An exciting application of our O-MAP platform that I hope will transform the study of nuclear architecture.
If you've ever wanted to dissect the subnuclear "neighborhood" around an individual locus, read on! (1/30)
I haven't seen a nucleic acids starter pack yet, so I started one! Please reach out if I missed you! go.bsky.app/NxKgUY5
Have you watched Twin Peaks?
More or less sense than using pipes as the primary means of transit?

Apparently since the beginning!
Thanks!
We designed the TECdisplay platform to be broadly accessible and easy to use, so our hope is that this approach democratizes the development and application of high-throughput RNA biochemical assays.

We applied this to a ZTP riboswitch variant library to investigate the mechanisms by which variants that match the consensus sequence and structure misfold. Panel c illustrates the ZMP-mediated antitermination response for the 32,768 variants in the library!

Our first TECdisplay assay measures transcription termination efficiency: TECs are tethered to a bead by the nascent RNA and DNA is released from the bead if transcription is terminated, but retained on the bead if RNA polymerase bypasses the terminator.
It also avoids the technical biases of RNA sequencing library preparation and is compatible with assays that destroy the RNA, since sequence information is recovered from DNA.
The activity of each RNA variant is then measured as the distribution of template DNA between fractions. In principle this approach can be applied to any RNA function that can be used to fractionate a TEC library.
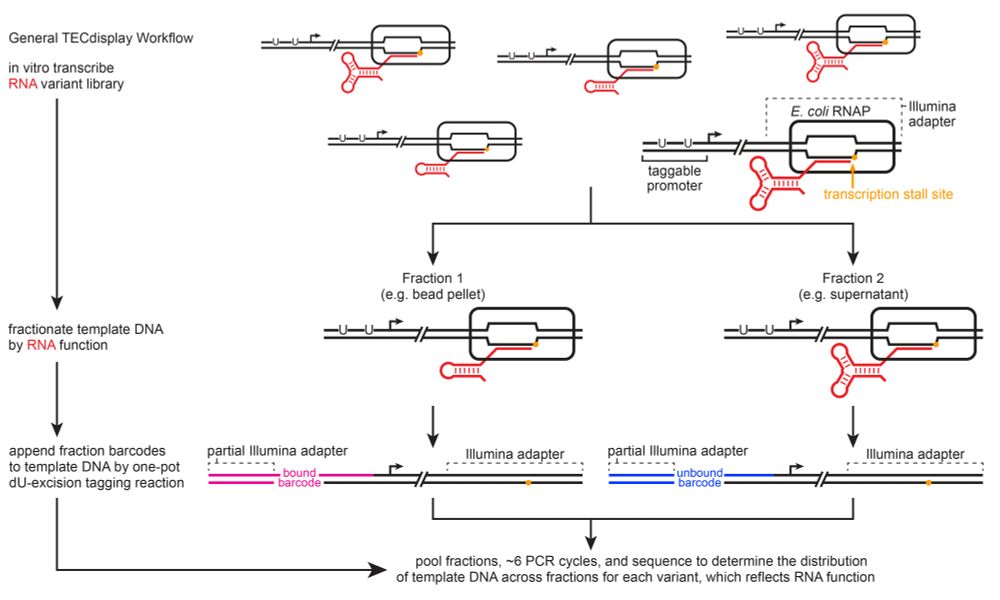
About 6 years ago, I had an idea for measuring RNA function by using the activity of cotransciptionally displayed RNA to fractionate a transcription elongation complex (TEC) library. After fractionation, DNA would be recovered, barcoded, pooled back together, and sequenced

Our latest preprint, led by Skyler Kelly: we developed a platform for high-throughput cotranscriptional RNA biochemistry called Transcription Elongation Complex display. www.biorxiv.org/content/10.1...
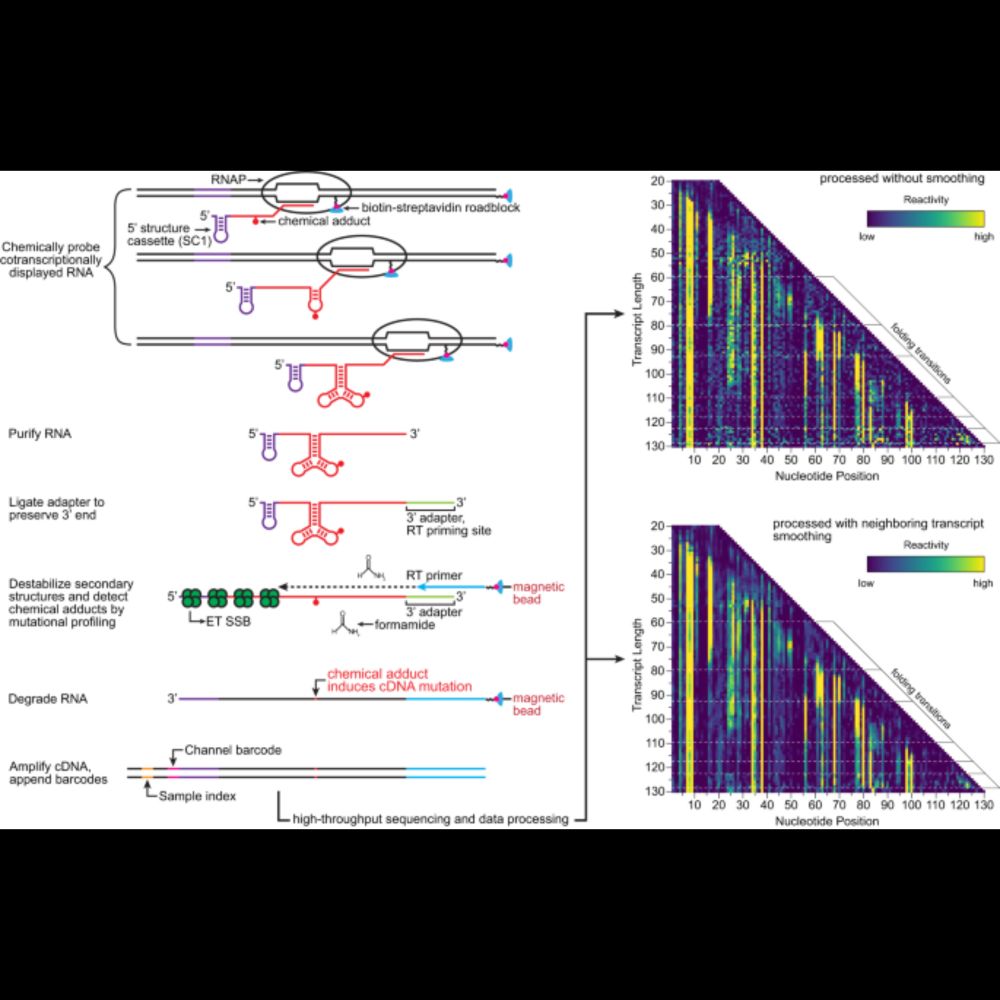
The final version of our article describing a modernized procedure for cotranscriptional RNA structure probing is online! www.nature.com/articles/s41...