Tvp complexes support formation of the VgrG-PAAR spike during Type VI secretion system assembly https://www.biorxiv.org/content/10.64898/2026.02.23.707444v1
Tvp complexes support formation of the VgrG-PAAR spike during Type VI secretion system assembly https://www.biorxiv.org/content/10.64898/2026.02.23.707444v1
Congratulations to MD-PhD student, Amanda Velez and all co-authors on our new PNAS paper describing how the innate immune protein calprotectin incapacitates autolysins, inducing tolerance to B-lactam antibiotics. A fun collaboration with Thomas Kehl-Fie. www.pnas.org/doi/10.1073/...

Congratulations to Sean Yu-Hao Wang, Will DePas, @catarmbruster.bsky.social and colleagues on their use of experimental evolution to identify regulators of Mycobacterium abscessus biofilm production!
www.sciencedirect.com/science/arti...

Regulatory Rewiring Drives Intraspecies Competition in Bacillus subtilis
bioRxiv from @bacteriacities.bsky.social
#competition #rapP #comP
www.biorxiv.org/content/10.1...

Schematic overview of the process of DNA-affinity / pull down identification of unknown nucleic acid-binding proteins
Identification of Previously Unknown DNA-Binding Proteins Using DNA Affinity/Pull-Down Methods Followed by Mass Spectrometry
Jutras, Babb, Jusufovic, Krusenstjerna, Saylor, Verma, and Stevenson
Current Protocols 2025, 5:e70264
doi: 10.1002/cpz1.70264
#MicroSky

🚨Preprint alert - this is a big one! We transfer the revolutionary power of TnSeq to bacteriophages.
Our HIDEN-SEQ links the "dark matter" genes of your favorite phage to any selectable phenotype, guiding the path from fun observations to molecular mechanisms.
A thread 1/8
Congratulations Alastair and Prerna! Exciting times :)
Congratulations, Jay! Looking forward to learning about your discoveries :)
An Rhs effector uses distinct target cell functions to intoxicate bacterial and fungal competitors https://www.biorxiv.org/content/10.1101/2025.10.28.685041v1
Independent research fellowships leading to tenured positions at the John Innes Centre.
Repost = nice. Thank you very much!!!
Cool!

New preprint from the lab! @findunmun1.bsky.social @chemsley.bsky.social We looked at new protein synthesis and genetic fitness during starvation in P. aeruginosa. Resources are dynamically redistributed to support ongoing activity, even over days of total starvation: www.biorxiv.org/content/10.1...
Congrats, Lev!
They will survive much better if you let them grow into stationary phase in rich media first, and then transfer them to a minimal medium with no carbon source. Transferring to carbon starvation directly from exponential growth causes faster death. See for example PMID: 32500952 and PMID: 34058747.
yes, the MMBs studied by george schaible and roland hatzenpichler are fascinating 𝗼𝗯𝗹𝗶𝗴𝗮𝘁𝗲 𝗺𝘂𝗹𝘁𝗶𝗰𝗲𝗹𝗹𝘂𝗹𝗮𝗿 bacteria 🤩
STC had mentioned them first in 2007 > schaechter.asmblog.org/schaechter/2...
and recently in > schaechter.asmblog.org/schaechter/2...
#MicroSky
In @molecularmicro.bsky.social: two special issues on Prokaryotic Chromosome Organization. Many excellent contributions, including a vision article: 'Future directions of the Prokaryotic Chromosome Field', outcome of BioPhyChrom 2023 @biophychrom.bsky.social onlinelibrary.wiley.com/doi/10.1111/...

Polyphosphate: The “Dark Matter” of Bacterial Chromatin Structure onlinelibrary.wiley.com/doi/10.1111/... #jcampubs
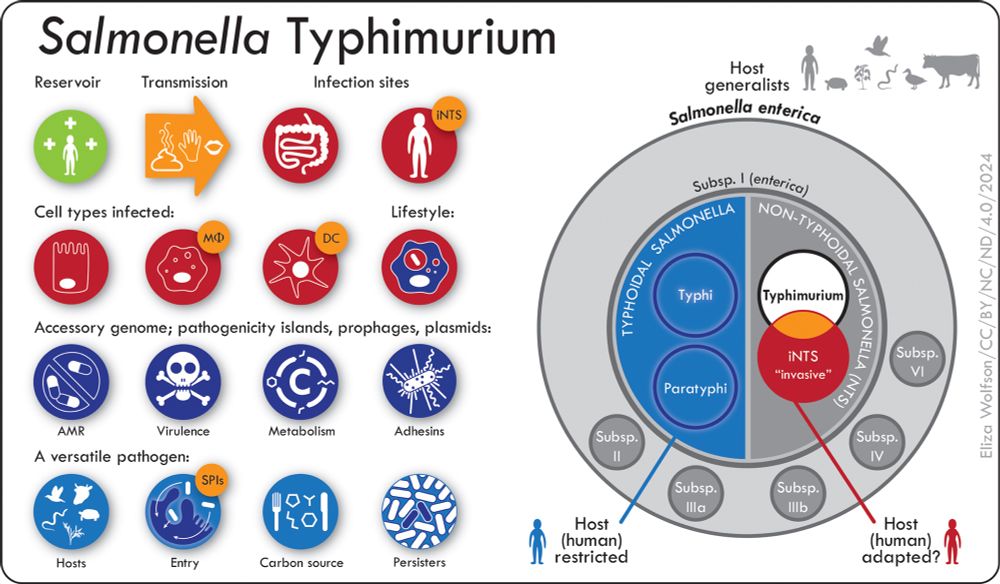
The taxonomy & lifestyle of Salmonella Typhimurium
www.microbiologyresearch.org/content/jour...

Congrats to Findlay and many thanks to examiners Dan Neill and Martin Welch for their work on a successful viva today! Findlay has worked so hard and accomplished so much while blazing the trail as the first PhD student in the lab :)
Together with @frimanscience.bsky.social and our Chinese colleagues, we have worked out how to predict
(i) siderophore molecular structures
www.biorxiv.org/content/10.1...
and
(ii) siderophore interaction networks in communities
www.biorxiv.org/content/10.1...
from sequence data!
Thanks for the introduction!