New preprint from the lab! We dig into the regulatory roles of TEs in the mouse placenta, with some surprising findings. Led by the awesome @smamante.bsky.social. Particular kudos to him for navigating the many twists and turns of the project.
New preprint from the lab! We dig into the regulatory roles of TEs in the mouse placenta, with some surprising findings. Led by the awesome @smamante.bsky.social. Particular kudos to him for navigating the many twists and turns of the project.

New preprint from our lab: ‘Retroviral insertions contributed to the divergence of human and chimpanzee brains’. Very proud of this piece which took almost a decade to finish.
www.biorxiv.org/content/10.6...

📣We are looking for a Project Assistant to join my group at the Wallenberg Center for Molecular Medicine at Umeå University. Join us if you are interested in transposable elements, epigenetics and human brain aging. 🧠🧬 Dont hesitate to reach out with any questions! 📣
umu.varbi.com/en/what:job/...

Had an absolute blast presenting my “Last Minute Breakthrough talk” at #EMBOmobilegenome today! 🔥 What an incredible crowd, the energy in the room was unreal. Huge thanks to the organizers for selecting me and to everyone who came, asked questions, and made it such a fun session! 🙌

Having a lot of fun at #EMBOMobileGenome 😃
Perfect timing for our paper from the lab of @toddmacfarlan.bsky.social to be out @natcomms.nature.com!!
…and I’m currently on the job market looking for a new scientific home!
www.nature.com/articles/s41...

How do pluripotent stem cells resist harmful interferon responses to safeguard development? Through total epigenetic lockdown of ligands, sensors and effectors of IFN-I. In our preprint, James Holt shares his PhD discoveries on the ground state of immune evasion 😊: www.biorxiv.org/cgi/content/...
🌟Im happy to share that this winter I'll be starting my lab at the Wallenberg Center for Molecular Medicine at Umeå University! Excited to join the Department of Medical and Translational Biology as a Wallenberg Fellow and looking forward to new collaborations and cool transposon research 🤩 🧬

The @crick.ac.uk is recruiting Early Career Group Leaders
- Lab set-up, research costs, salaries for up to 5 researchers
- Support for up to 12 years
- Access to our core facilities
- Competitive salary
- Fantastic colleagues
- All areas of biology
Deadline 27 Nov
www.crick.ac.uk/careers-stud...
⚡⚡Excited to announce I'll be starting my lab at the Max Planck Institute of Molecular Genetics (@molgen.mpg.de) in Berlin in December! Leaving sunny California to join a fantastic environment with colleagues who do super cool work.
🔬🦠I'm hiring at all levels! 🔬🦠Check: www.molgen.mpg.de/fueyo-lab
Well done!!!

#1 Centromeres are epigenetic loci defined by CENP-A, positioned in unmethylated DNA flanked by highly methylated regions. Our work, published in @natgenet.nature.com in collaboration with @naltemose.bsky.social investigates the role of DNAme at human centromeres www.nature.com/articles/s41...
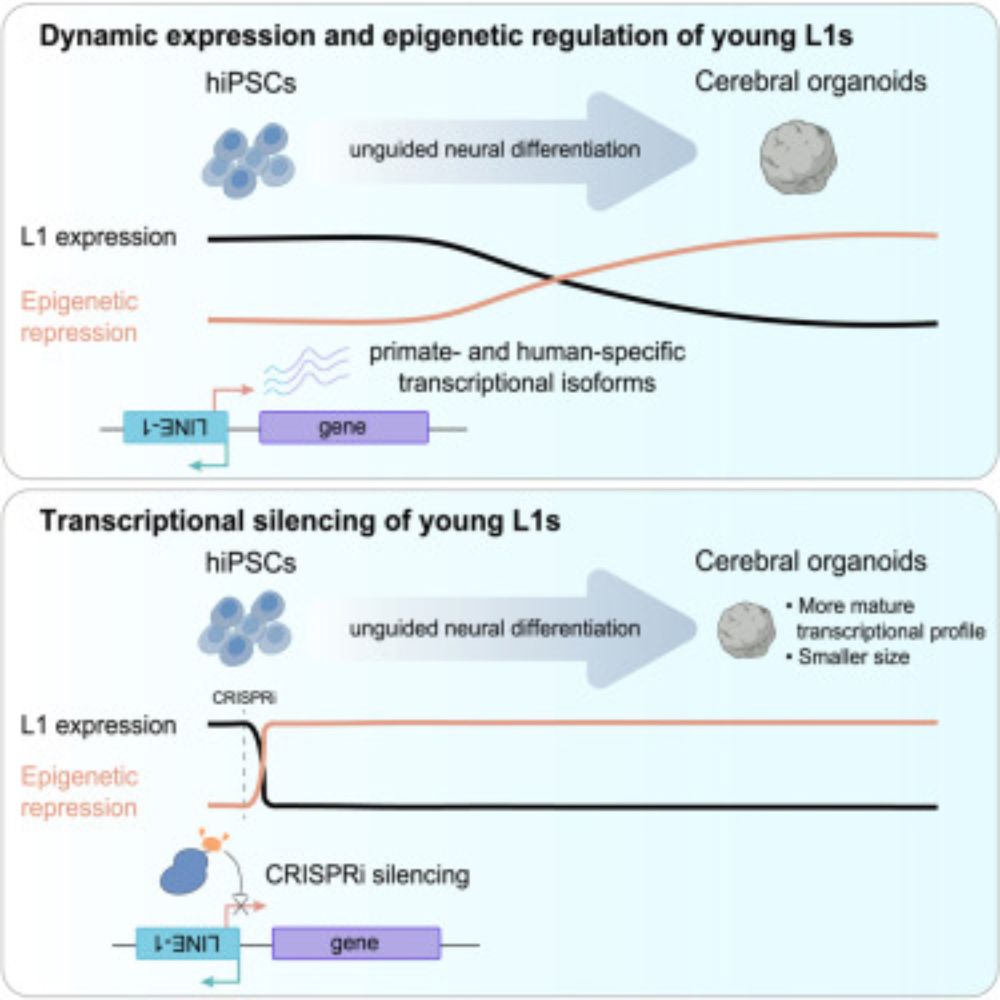
New paper from our lab! We found that LINE-1 transposons contribute to early human brain development.
Funded by @asapresearch.parkinsonsroadmap.org
www.cell.com/cell-genomic...
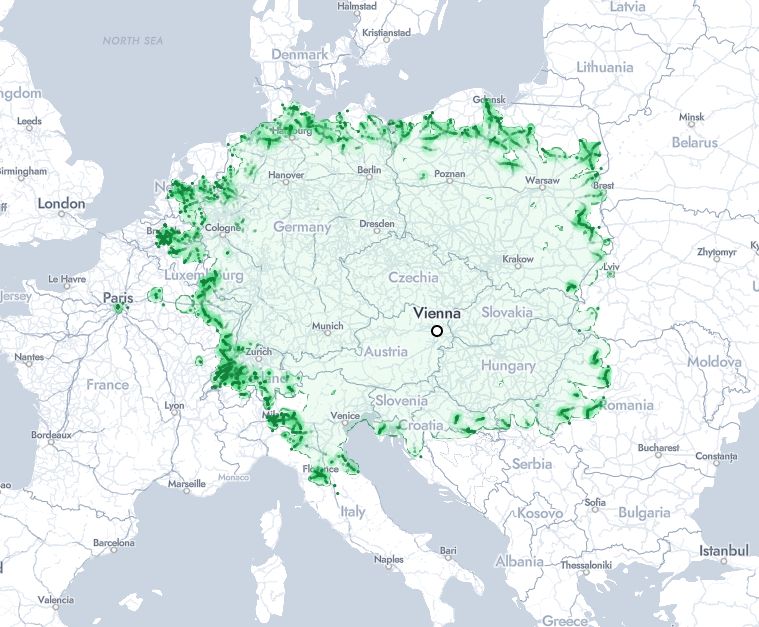
Cities reachable by train from Vienna in under 10 hours
Civilization, visualized
Positively mesmerizing: www.chronotrains.com/en/explore/2...

Nature research paper: Complex genetic variation in nearly complete human genomes
go.nature.com/44YuyIV

Very happy to share our protocols paper for CELLO-seq. This will make single cell long read RNA-seq more accessible and provides analysis guidelines. We hope this helps the #transposon #TEsky community and folks working on #singleCell isoform and allelic #gene expression. doi.org/10.1038/s415...
A must read!
We are looking for new colleagues to come join us in Galway as group leaders (Junior and Senior). The Centre for Chromosome Biology is a great place and it is a good time to join. Please reach out if you want to chat about the opportunity!
www.nature.com/naturecareer...

Finally out! 🥳 Our paper showing how a transposable element (TE) insertion can cause developmental phenotypes is now published @natgenet.nature.com 🧬🦠🐁
Below is a brief description of the major findings. Check the full version of the paper for more details: www.nature.com/articles/s41588-025-02248-5
The work was led by my inspiring PhD student Fereshteh Dorazehi and ably supported by a team of wonderful scientists over many years! Fereshteh and I have presented/will present the story at some recent and upcoming meetings and are happy to hear feedback. For now it's time for summer holidays! 9/9
Second, the factor directing human L1 methylation in early development was unknown - a deletion in the ancestral promoter is thought to evade silencing by the KRAB-ZNF/TRIM28 system. We show MORC2 is critical here, at least for those most dangerous elements transcribed in the pluripotent state. 8/X
First, MORC2 ATPase mutations cause severe developmental disorders: we explain how ATPase activity is coupled to chromatin binding and CpG methylation patterning of different targets. That many of the hyperrepressed genes encode KRAB-ZNFs, themselves TE silencers, is conceptually fascinating! 7/X
Apart from relating human genetic L1 activation to a clear cellular phenotype in development cell models, we are excited about these results for two main reasons...
6/X
Genetic loss of L1 silencing due to MORC2 mutation had severe consequences on cellular fitness in a 2D NPC model after pluripotency exit - rescued upon L1 CRISPRi! (shout out to @anitaada.bsky.social @jakobssonlab.bsky.social for the vector - see www.biorxiv.org/content/10.1... ) 5/X
Explaining the persistence of these transcriptional phenotypes we found striking loss of CpG methylation over the youngest human- and hominoid specific L1s (including polymorphic alleles), and hypermethylation of gene promoters. 4/X
Modelling these effects in pluripotent stem cells, we observed loss of function (transcriptional activation) at L1s and gain of function (transcriptional repression) at promoters, an effect which persisted upon differentiation to neural progenitors. 3/X
We engineered mutations to figure out requirements of MORC2 binding to its distinct chromatin targets. Accumulation of MORC2 at L1s and other broad targets requires reversible ATP-dependent dimerization, whereas engagement of gene promoters does not. Mutation misdirects MORC2 onto promoters. 2/X

Very proud of this work where we define how the MORC2 ATPase directs CpG methylation of active human L1 transposons in early development.
Thanks to co-authors, funders & wonderful environment @lundstem.bsky.social
www.biorxiv.org/content/10.1...
Brief thread below (do people still do that?) 1/X

We wrote a review on Transposable Elements (TEs) and almost all aspects of TE silencing and their roles in biological processes & disease.
www.nature.com/articles/s41...
A major KI initiative to recruit new assistant professors with outstanding proposals in all areas of medicine, biomedicine and public health. We offer an amazing research environment, great colleagues and generous startup packages. Check it out and get working on your applications! (repost please!)
Lund University is offering 2-year visiting professorships (20% time) for full professors, with a deadline of August 17. Women are encouraged to apply. More info: https://lu.varbi.com/en/what:job/jobID:835541/ #job