Looks cool, thank you! Would it also work for conventional short-read (high depth) scRNAseq - with fewer variants detected due to low transcript coverage or are there fundamental differences to long-read that make this harder?
Looks cool, thank you! Would it also work for conventional short-read (high depth) scRNAseq - with fewer variants detected due to low transcript coverage or are there fundamental differences to long-read that make this harder?
Sounds exciting! What approach are you using to identify and call the variants confidently?



📣 SIC Summer School 2026 — applications open!
📍Hornbæk, DK | 18–20 May 2026
🎯Theme: Skin Immunity & Homeostasis
💵Free (stay + meals). For PhD students & postdocs.
✍️Deadline: 15 Jan 2026 (23:59 CET).
🔗Full details: sic.ku.dk/summerschool/
🔗Register here: eventsignup.ku.dk/sicss2026
Glad to see our article on how cancer drugs and bacterial (S. aureus) infections can affect co-existing malignant subclones very differently in cutaneous T-cell lymphoma.
Thank you Cancer Discovery (@aacrjournals.bsky.social) for the beautiful title-page graphics!
Congratulations Terkild and the team👏💪😊 very important new knowledge on the intricacies of this horrible cancer - this new understanding of coexisting yet specialized cancer subclones is a key discovery for a truly targeted treatment in the future !!!!
Bravo 👏👏👏

Finally, we would also like to thank N. Borcherding (@thehumanborch.bsky.social), M. Khodadoust, A. Herrera and A. Cheng for making their data public and their gracious assistance in the annotation and elaboration of their respective single-cell RNA sequencing data sets.
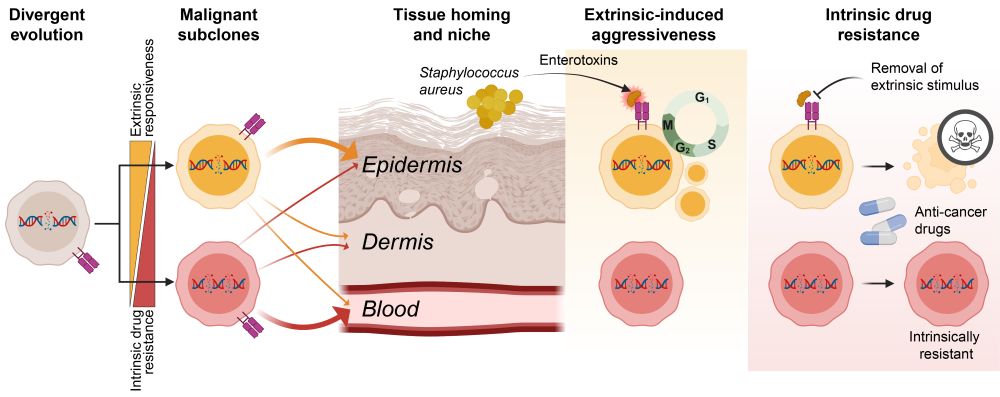
Schematic figure showing divergent evolution of malignant subclones and how they migrate to different tissue compartments (blood vs skin), respond differently to extrinsic stimuli such as Staphylococcus aureus enterotoxins, and how subclones differ in their intrinsic resistance to anti-cancer drugs.
❗How can a cancer exploit its environment and still resist treatment?
✅The answer: co-existing malignant subclones.
Let me walk you through our latest study investigating how divergent evolution drives adaptability, aggressiveness, and drug resistance of T cell cancer. 1/🧵
doi.org/10.1158/2159...

We greatly appreciate the constructive and professional feedback and suggestions from our reviewers and our editor @elizsmckenna.bsky.social at Cancer Discovery @theaacr.bsky.social!
We also thank the LEO foundation and the Danish Cancer Society @cancer.dk for funding the project!
This work was a team effort involving so many fantastic collaborators, clinicians, patients and their loved ones - and of course the incredible people in the group of @niels-odum.bsky.social
Too many people involved to list here - so go check out the paper!
doi.org/10.1158/2159...
🎧 Prefer audio? We’ve generated two AI-narrated summaries of the study:
– A brief summary (~15 min) tinyurl.com/bdh6h9tb
– A more in-depth version (~45 min) tinyurl.com/4nunr2v4
Great for listening on the go while diving into the details. 10/

Our findings underscore that understanding this complex network of specialized cancer subclones is key to overcoming treatment resistance in L-CTCL. Mapping a patient's subclonal landscape could guide personalized, rational combination therapies targeting all subclones for enduring response. 9/

This discovery offers an explanation for varying treatment responses across different compartments (skin/blood) in L-CTCL patients. By targeting bacterial infections and dampening inflammation, we expose inherent vulnerabilities, making otherwise hard-to-treat subclones susceptible to therapy. 8/

Schematic figure showing divergent evolution of malignant subclones and how they migrate to different tissue compartments (blood vs skin), respond differently to extrinsic stimuli such as Staphylococcus aureus enterotoxins, and how subclones differ in their intrinsic resistance to anti-cancer drugs.
Divergent evolution results in co-existing subclones that have different tissue preferences and respond differently to external factors: cytokines and bacterial infections. The most aggressive subclones are also most sensitive to treatment if stimuli are removed exposing their vulnerabilities. 7/

Figure showing marked difference in the degree a clinically relevant anti-cancer drug (HDAC-inhibitor) induced cell death between divergent malignant subclones from a L-CTCL patient. Similar analyses across 6 different anti-cancer drugs (Vorinostat, Romidepsin, Etoposide, Oligomycin, Doxorubicin and Bortezomib) is summarize for subclones across 12 L-CTCL patients by flow cytometry. Also shows schematic conclusion figure showin that the subclones that are most responsive to extrinsic stimuli such as Staphylococcus aureus enterotoxins (SE) are also the most sensitive to anti-cancer drugs - but only if the stimuli are removed. Other subclones that do not respond to extrinsic stimuli are intrinsically more drug-resistant.
Here's the trade-off ⚖️: The subclones that respond most strongly to 🦠S. aureus toxins—the aggressive ones—are also the most intrinsically sensitive to 💊anti-cancer drugs when the stimuli are removed.
This means reducing inflammation could unmask therapeutic vulnerabilities in aggressive clones. 6/

Figure showing differential activation of two malignant subclone from a L-CTCL patient following exposure to Staphylococcus aureus supernatants. Similar analysis across subclones from 16 L-CTCL patients is shown which highlights that divergent subclones can be divided into four distinct Staphylococcus aureus enterotoxin (SE) response groups and that the subclones assigned to group 3 all exhibit higher levels of proliferation than their neighboring subclones when cultured in the presence of SE. Final schematic is summarizing this by showing that divergent subclones respond differential to enterotoxins leading to increased activation, transcription and proliferation of responsive subclones.
Bacterial (S. aureus) infections are a major clinical problem in CTCL, fueling malignant growth and persistence. S. aureus toxins selectively activate certain subclones, while leaving others largely unaffected. This provides a selective advantage to responsive subclones!
But it comes at a price… 5/

Plots and schematic showing experimental setup for matched blood and skin (separated into dermis and epidermis for further spatial detail). Barplots showing subclone abundance across three patients across blood, dermis and epidermis from two independent lesional sites. Also includes a schematic figure showing that malignant subclones exhibit differences in the tissue preference - some prefer blood, others prefer dermis or epidermis. Final plot shows correlation between subclones’ ability to respond to their (inflammatory) environment and their proliferation (indicating aggressiveness) in the skin. Schematic showing that the subclone most abundant in blood (shown in red) tends to have lower levels of interaction with the microenvironment in the skin - leading to a growth advantage of the responsive subclones.
These subclones show differences in tissue homing (some prefer blood, others skin), metabolism, and immune signaling. Intriguingly, the subclone most abundant in blood often has less capacity for interacting with their inflammatory environment, correlating with reduced proliferation in the skin. 4/

Schematic figure showing experimental setup with collection of matched blood and skin samples from 57 leukemic cutaneous T-cell lymphoma (L-CTCL) patients as well as the multimodal single-cell analysis approach combining single-cell T-cell receptor (scTCR) and cellular indexing of transcriptomes and epitopes (CITE) sequencing: scTCR+CITE-seq. Also showing data from each of the 57 included L-CTCL patients showing their relative abundance of different subclones as percentage of total CD4+ T cell population. This plot shows that distinct malignant subclones could be identified in more than 80% of the L-CTCL patients. Final plot shows that subclones co-exist over extended periods of time by a connected barplot with subclone abundance at multiple timepoints up to 294 days.
Using advanced single-cell analysis (scTCR+CITE-seq) on matched skin and blood, we found 🧬genetically distinct malignant subclones in more than 80% of L-CTCL patients. Subclones have distinct RNA+protein signatures, are shared between blood and skin, and co-exist over extended periods of time⏳. 3/

Schematic figure showing experimental setup with collection of matched blood and skin samples from 57 leukemic cutaneous T-cell lymphoma (L-CTCL) patients as well as the multimodal single-cell analysis approach combining single-cell T-cell receptor (scTCR) and cellular indexing of transcriptomes and epitopes (CITE) sequencing: scTCR+CITE-seq. Figure shows the five main findings of the study: 1. Divergent evolution resulting in multiple subclones with distinct extrinsic responsiveness and intrinsic drug resistance, 2. Subclones are functionally and genomically distinct, 3. Subclones home to different tissue niches (some prefer blood whereas others prefer the dermis or epidermis of the skin), 4. Divergent subclones exhibit differences in their induced aggressiveness visualized by different response to Staphylococcus aureus enterotoxins, and 5. Divergent subclones exhibit differences in their intrinsic drug resistance. Aggressive subclones become sensitive to treatment when their extrinsic stimuli are removed.
TL;DR
👉Leukemic CTCL patients harbor functionally distinct co-existing subclones
👉Subclones respond differently to external factors such as cytokines, bacterial infections, and cancer drugs
👉The most aggressive subclones are also most sensitive to treatment if their extrinsic stimuli are removed
2/

Schematic figure showing divergent evolution of malignant subclones and how they migrate to different tissue compartments (blood vs skin), respond differently to extrinsic stimuli such as Staphylococcus aureus enterotoxins, and how subclones differ in their intrinsic resistance to anti-cancer drugs.
❗How can a cancer exploit its environment and still resist treatment?
✅The answer: co-existing malignant subclones.
Let me walk you through our latest study investigating how divergent evolution drives adaptability, aggressiveness, and drug resistance of T cell cancer. 1/🧵
doi.org/10.1158/2159...
🎧 Prefer audio? We’ve generated two AI-narrated summaries of the study:
– A brief summary (~15 min) tinyurl.com/bdh6h9tb
– A more in-depth version (~45 min) tinyurl.com/4nunr2v4
Great for listening on the go while diving into the details. 10/

Our findings underscore that understanding this complex network of specialized cancer subclones is key to overcoming treatment resistance in L-CTCL. Mapping a patient's subclonal landscape could guide personalized, rational combination therapies targeting all subclones for enduring response. 9/

This discovery offers an explanation for varying treatment responses across different compartments (skin/blood) in L-CTCL patients. By targeting bacterial infections and dampening inflammation, we expose these inherent vulnerabilities, making hard-to-treat subclones susceptible to therapy. 8/

Schematic figure showing divergent evolution of malignant subclones and how they migrate to different tissue compartments (blood vs skin), respond differently to extrinsic stimuli such as Staphylococcus aureus enterotoxins, and how subclones differ in their intrinsic resistance to anti-cancer drugs.
Divergent evolution results in co-existing subclones that have different tissue preferences and respond differently to external factors such as cytokines and infections. The most aggressive subclones are also most sensitive to treatment if stimuli are removed exposing their vulnerabilities. 7/

Figure showing marked difference in the degree a clinically relevant anti-cancer drug (HDAC-inhibitor) induced cell death between divergent malignant subclones from a L-CTCL patient. Similar analyses across 6 different anti-cancer drugs (Vorinostat, Romidepsin, Etoposide, Oligomycin, Doxorubicin and Bortezomib) is summarize for subclones across 12 L-CTCL patients by flow cytometry. Also shows schematic conclusion figure showin that the subclones that are most responsive to extrinsic stimuli such as Staphylococcus aureus enterotoxins (SE) are also the most sensitive to anti-cancer drugs - but only if the stimuli are removed. Other subclones that do not respond to extrinsic stimuli are intrinsically more drug-resistant.
Here's the trade-off ⚖️: The subclones that respond most strongly to 🦠S. aureus toxins—the aggressive ones—are also the most intrinsically sensitive to 💊anti-cancer drugs when the stimuli are removed.
This means reducing inflammation could unmask therapeutic vulnerabilities in aggressive clones. 6/

Figure showing differential activation of two malignant subclone from a L-CTCL patient following exposure to Staphylococcus aureus supernatants. Similar analysis across subclones from 16 L-CTCL patients is shown which highlights that divergent subclones can be divided into four distinct Staphylococcus aureus enterotoxin (SE) response groups and that the subclones assigned to group 3 all exhibit higher levels of proliferation than their neighboring subclones when cultured in the presence of SE. Final schematic is summarizing this by showing that divergent subclones respond differential to enterotoxins leading to increased activation, transcription and proliferation of responsive subclones.
Bacterial (S. aureus) infections are a major clinical problem in CTCL, fueling malignant growth and persistence. S. aureus toxins selectively activate certain subclones, while leaving others largely unaffected. This provides a selective advantage to responsive subclones!
But it comes at a price… 5/

Plots and schematic showing experimental setup for matched blood and skin (separated into dermis and epidermis for further spatial detail). Barplots showing subclone abundance across three patients across blood, dermis and epidermis from two independent lesional sites. Also includes a schematic figure showing that malignant subclones exhibit differences in the tissue preference - some prefer blood, others prefer dermis or epidermis. Final plot shows correlation between subclones’ ability to respond to their (inflammatory) environment and their proliferation (indicating aggressiveness) in the skin. Schematic showing that the subclone most abundant in blood (shown in red) tends to have lower levels of interaction with the microenvironment in the skin - leading to a growth advantage of the responsive subclones.
These subclones show differences in tissue homing (some prefer blood, others skin), metabolism, and immune signaling. Intriguingly, the subclone most abundant in blood often has less capacity for interacting with their inflammatory environment, correlating with reduced proliferation in the skin. 4/

Schematic figure showing experimental setup with collection of matched blood and skin samples from 57 leukemic cutaneous T-cell lymphoma (L-CTCL) patients as well as the multimodal single-cell analysis approach combining single-cell T-cell receptor (scTCR) and cellular indexing of transcriptomes and epitopes (CITE) sequencing: scTCR+CITE-seq. Also showing data from each of the 57 included L-CTCL patients showing their relative abundance of different subclones as percentage of total CD4+ T cell population. This plot shows that distinct malignant subclones could be identified in more than 80% of the L-CTCL patients. Final plot shows that subclones co-exist over extended periods of time by a connected barplot with subclone abundance at multiple timepoints up to 294 days.
Using advanced single-cell analysis (scTCR+CITE-seq) on matched skin and blood, we found 🧬genetically distinct malignant subclones in more than 80% of L-CTCL patients. Subclones have distinct RNA+protein signatures, are shared between blood and skin, and co-exist over extended periods of time⏳. 3/

Schematic figure showing experimental setup with collection of matched blood and skin samples from 57 leukemic cutaneous T-cell lymphoma (L-CTCL) patients as well as the multimodal single-cell analysis approach combining single-cell T-cell receptor (scTCR) and cellular indexing of transcriptomes and epitopes (CITE) sequencing: scTCR+CITE-seq. Figure shows the five main findings of the study: 1. Divergent evolution resulting in multiple subclones with distinct extrinsic responsiveness and intrinsic drug resistance, 2. Subclones are functionally and genomically distinct, 3. Subclones home to different tissue niches (some prefer blood whereas others prefer the dermis or epidermis of the skin), 4. Divergent subclones exhibit differences in their induced aggressiveness visualized by different response to Staphylococcus aureus enterotoxins, and 5. Divergent subclones exhibit differences in their intrinsic drug resistance. Aggressive subclones become sensitive to treatment when their extrinsic stimuli are removed.
TL;DR
👉Leukemic CTCL patients harbor functionally distinct co-existing subclones
👉Subclones respond differently to external factors such as cytokines, bacterial infections, and cancer drugs
👉The most aggressive subclones are also most sensitive to treatment if their extrinsic stimuli are removed
2/

Now online in Cancer Discovery: Divergent Evolution of Malignant Subclones Maintains a Balance Between Induced Aggressiveness and Intrinsic Drug Resistance in T Cell Cancer - by @terkild.bsky.social, @niels-odum.bsky.social, and colleagues doi.org/10.1158/2159... @ucph.bsky.social
Subclonal, tumor microenvironment and bacterial interactions in cutaneous lymphoma - particularly how bacteria fuels the cancer.