We compared 3 MS and 1 aptamer proteomics workflows on elite athlete plasma. Platform differences aren't just technical—they reveal distinct biology of exercise adaptation and metabolic health. New preprint just in time for Winter Olympics 🏆 🥇
biorxiv.org/content/10.6...
09.02.2026 08:10
👍 5
🔁 1
💬 0
📌 1

An adaptive, continuous-learning framework for clinical decision-making from proteome-wide biofluid data www.nature.com/artic...
---
#proteomics #prot-paper
28.01.2026 10:00
👍 4
🔁 1
💬 0
📌 0

Bead-based plasma #proteomics workflows boost proteome coverage but are vulnerable to cellular contamination, highlighting the need for contamination-aware quality control in #biomarker discovery.
K. Korff, M. Mann, P. Geyer & colleagues @mpibiochem.bsky.social
🗞️ doi.org/10.1038/s443...
16.09.2025 08:54
👍 3
🔁 3
💬 0
📌 0

Pre-analytical drivers of bias in bead-enriched plasma proteomics | EMBO Molecular Medicine www.embopress.org/do...
---
#proteomics #prot-paper
13.09.2025 08:40
👍 13
🔁 5
💬 2
📌 0

Ontology-guided clustering enables #Proteomic analysis of rare #PediatricDisorders
By Ericka CM Itang, Johannes Mueller-Reif and colleagues @mpibiochem.bsky.social & German Center for Child and Adolescent Health
🗞️ doi.org/10.1038/s443...
27.05.2025 19:22
👍 3
🔁 1
💬 0
📌 0

Our new paper is out in EMBO Mol Med: we combine clinical ontologies with deep proteomics to study rare pediatric diseases. A step toward scalable, systems-level insight into child health.
Grateful to all collaborators & families.
www.embopress.org/doi/full/10....
27.05.2025 12:23
👍 0
🔁 0
💬 0
📌 0
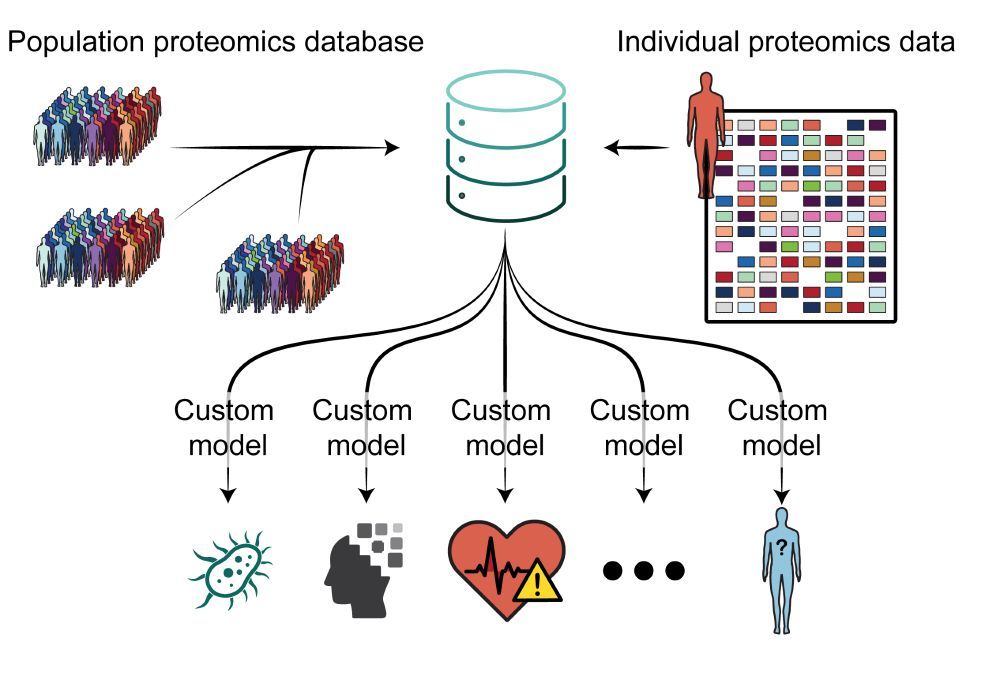
Check out ADAPT-MS: our flexible, scalable diagnostic framework that transforms discovery plasma proteomics directly into clinical decisions.Dynamic retraining enables single-sample diagnostics for any clinical question.
www.medrxiv.org/content/10.1...
06.05.2025 07:27
👍 12
🔁 6
💬 1
📌 0
In summary we propose to use discovery proteomics data directly for diagnostic and in future prognostic procedures, bypassing extensive assay development. ADAPT-MS is the framework we developed to enable this vision.
05.05.2025 19:37
👍 0
🔁 0
💬 0
📌 0
The beauty is that with every measured sample, the prior knowledge growth and enables better and more classification tasks. With growing databases these can be tailored to ever more specific subpopulations fitted to covariates and local specific effects.
05.05.2025 19:37
👍 0
🔁 0
💬 1
📌 0
This procedure again brought up wider implications: if we can classify one diagnostic question from discovery proteomics data, we can do this for all. The precondition is prior data for this question. With recent population based proteomics efforts this seems to be in reach in not to distant future.
05.05.2025 19:37
👍 0
🔁 0
💬 1
📌 0
Based on the overlap of this feature list and the available proteins per sample we train sample specific classifiers for each later measured sample where this decision is needed for. This makes the procedure robust and performant.
05.05.2025 19:37
👍 0
🔁 0
💬 1
📌 0
So we set out to build an ML architecture that can be employed to single sample discovery data without the need to process the sample within a study. The solution for us was a refitting procedure: we preselect features from study data for a specific task, e.g. benign from malignant classification.
05.05.2025 19:37
👍 0
🔁 0
💬 1
📌 0
After measuring a recent study and building successful classifiers with ML we asked ourselves: could we measure another patient sample now or in half a year and classify with the same performance? Admittedly, the answer is: it is complicated.
05.05.2025 19:37
👍 0
🔁 0
💬 1
📌 0

A regular question from clinical partners to me is: can you tell me if this patient has disease A or B from the plasma proteome? This sparked a new idea: ADAPT-MS is a concept framework enabling diagnostic classification from single sample discovery proteomics data.
www.medrxiv.org/content/10.1...
05.05.2025 19:37
👍 2
🔁 0
💬 1
📌 0
An adaptive, continuous-learning framework for clinical decision-making from proteome-wide biofluid data https://www.medrxiv.org/content/10.1101/2025.05.02.25326901v1
05.05.2025 15:41
👍 1
🔁 1
💬 0
📌 0
A simplified perchloric acid workflow with neutralization (PCA-N) for democratizing deep plasma proteomics at population scale https://www.biorxiv.org/content/10.1101/2025.03.24.645089v1
26.03.2025 23:45
👍 1
🔁 1
💬 0
📌 1
A simplified perchloric acid workflow with neutralization (PCA-N) for democratizing deep plasma proteomics at population scale https://www.biorxiv.org/content/10.1101/2025.03.24.645089v1
26.03.2025 23:45
👍 2
🔁 1
💬 0
📌 0








