www.nature.com/articles/d41...
www.nature.com/articles/d41...



🦠🔬🤖🧑💻 #mAIcrobe is out! With @pinholab.bsky.social's lab, we launched an open-source framework for high-throughput bacterial image analysis. By rockstars A. Brito & B. Saraiva et al, making #DeepLearning for phenotyping accessible! Easy to use, plus model training
📜 www.biorxiv.org/content/10.1...
🚀🔬 Announcing #IMS2026 at @itqbnova.bsky.social! We're assembling a #LifeSciences #Microscopy symposium packed with ✨ speakers and 🧑🔬 workshops. All about imaging cells with photons, electrons and AI! If you love microscopy, you need to be here. Join us on March 19!
ims2026.itqbnovacommunity.org
Congratulations!! 🥳🥳🥳

Our new preprint is up! This is the main postdoc work of @wiesner-t.bsky.social focusing on exocytosis along the axon shaft and its regulation by the sub membrane actin-spectrin scaffold: www.biorxiv.org/content/10.1...
Read the thread below for a summary of our findings 🧵1/11
Thank you @smlm-symposium.bsky.social for the opportunity to present at #SMLMS2025! 🎉 Stay tuned for the upcoming preprint! 📄
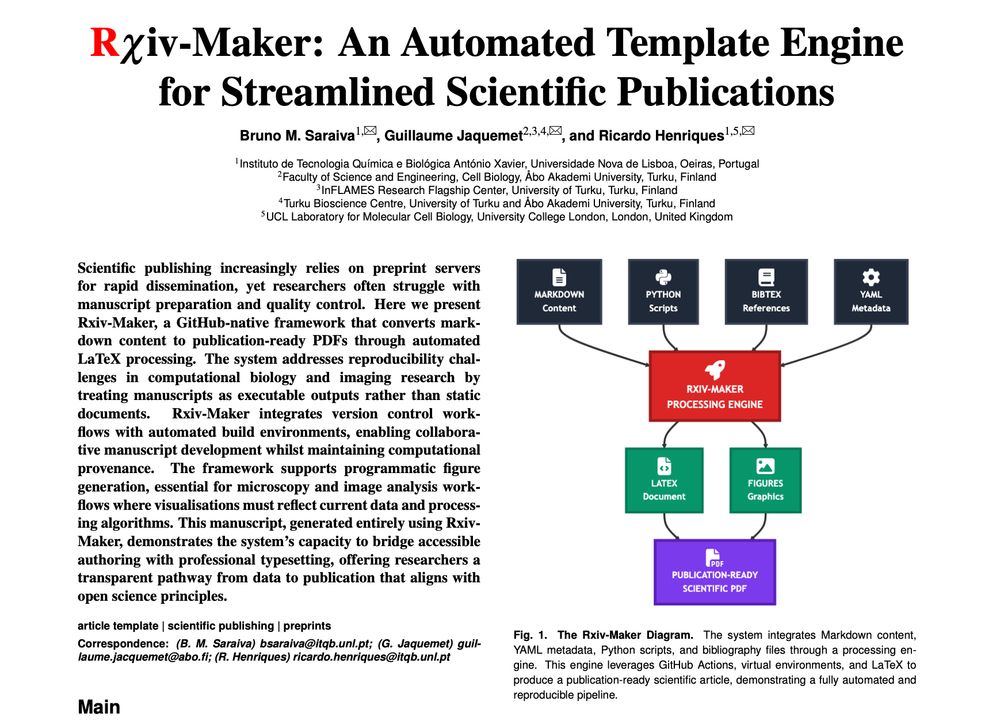



🧑🔬📰🚀 #RxivMaker is here! A framework automating the creation of publication-ready scientific #preprints from simple Markdown. Goodbye manual formatting, and hello gorgeous manuscripts! A great adventure with @guijacquemet.bsky.social's lab
📜: zenodo.org/records/1575...
👨💻: github.com/HenriquesLab...
Are you looking for a postdoc?
I will be looking for 2-4 people after the summer to join the lab !!
Do not hesitate to reach out and disseminate !!
AI4CellFate: Interpretable Early Cell Fate Prediction with Generative AI https://www.biorxiv.org/content/10.1101/2025.05.12.653464v1
And a big thank you to Dr. Alix Le Marois and Dr. Erik Sahai for providing this amazing dataset❤️(see "Imaging of MAP kinase dynamics reveals endocytic regulation of pulsatile signalling and network re-wiring in response to targeted therapy in EGFR-mutant non-small cell lung cancer")

A huge thank you ❤️ to @lucapanconi.bsky.social , @sebjbauer.bsky.social , Emma Latron, Maxime Gestin, Erik Sahai, Alix Le Marois, @juliettegriffie.bsky.social and our funders @cancerresearchuk.org , @kawresearch.bsky.social , @erc.europa.eu
biorxiv.org/content/10.1... (5/5)
Our model is small and versatile – this allows us to train it on a common laptop on the whole dataset in under 20 minutes ⚡️😌and use limited manual data annotations✍️ for accurate predictions (4/n)
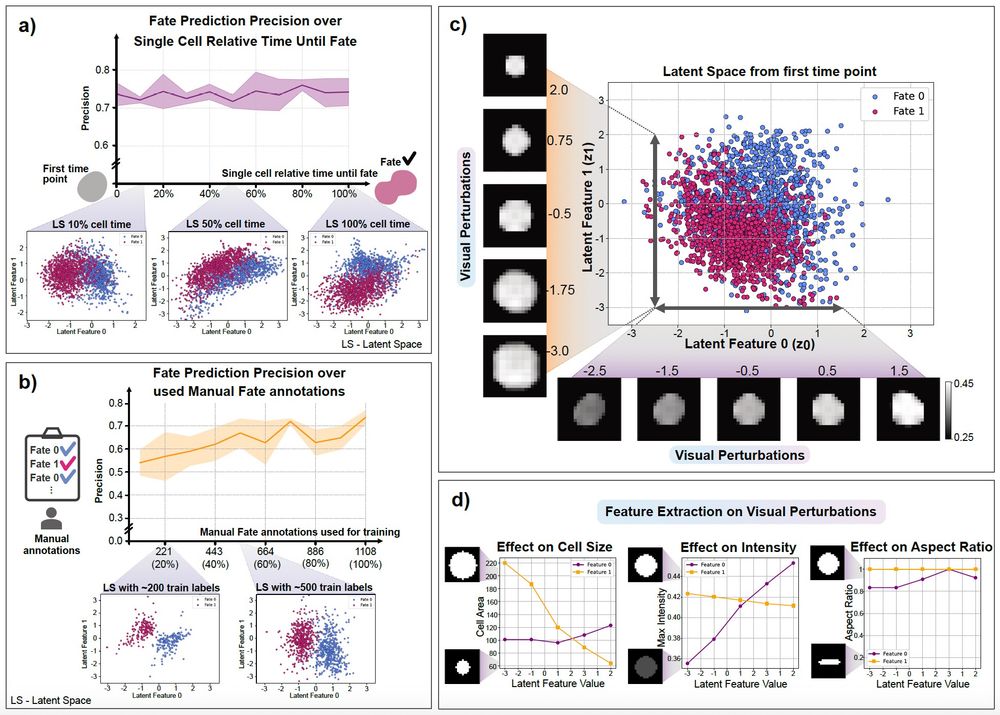
We were able to reliably predict cell fate from the very first frame ⚡️of 3 days long acquisition, and identify relevant biological processes🧬 involved in cancer persister cells, despite their known biological variability (3/n)
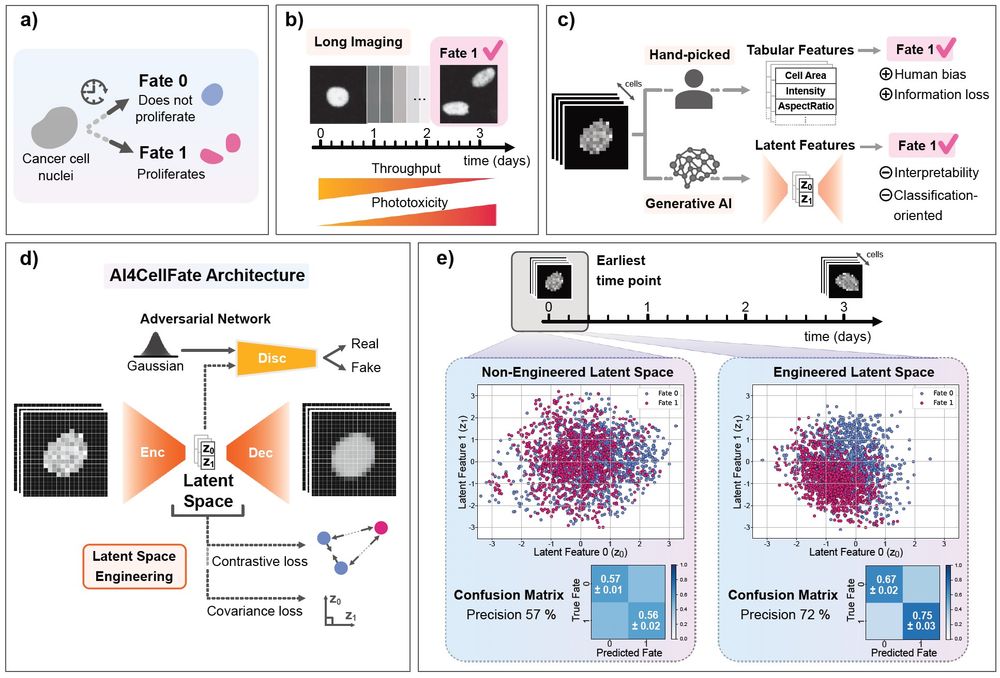
We propose a fully data-driven approach to early cell fate prediction from a microscopy time-lapse. We combine an adversarial autoencoder architecture with a multi-objective loss function, including contrastive learning, to optimise both cell fate prediction and interpretability (2/n)

Our work “AI4CellFate: Interpretable Early Cell Fate Prediction with Generative AI” is finally out in
@biorxivpreprint.bsky.social !!! 😍❤️ biorxiv.org/content/10.1... @scilifelab.se , @crick.ac.uk . More about it in the thread #AI4CellFate (1/n)⬇️


First research manuscript of my PhD finally submitted!! 🥳❤️ Preprint coming soon... 👀 @juliettegriffie.bsky.social
Come work with us! 😜 💻🔬

#NanoPyx and #LiquidEngine hitting the national news, highlighting our research @itqbnova.bsky.social
www.rtp.pt/noticias/pai...
📰: www.nature.com/articles/s41...
What an incredible journey with Bruno and António (not on bsky yet) ❤️ I learned so much with these guys and had a blast working with them. This was truly the dream team ❤️ #NanoPyx
Video 4: "How to Implement your Liquid Engine Method" youtu.be/gRGEjdT8opY (8/8)

Video 3: "How to Benchmark your Implementations with the Liquid Engine in 1 minute" (7/n) youtu.be/9hF7nLtFzoo

Video 2: "How to Build your Liquid Engine Class in 1 minute" youtu.be/QQsXrZ_jFa8 (6/n)

HOW CAN YOU USE #NanoPyx ? We recorded a series of 1-minute video tutorials that showcase how you can use #NanoPyx, and the capabilities of the #LiquidEngine on your own image analysis workflows! Video 1: youtu.be/Dx2lHoRB044 (5/n)
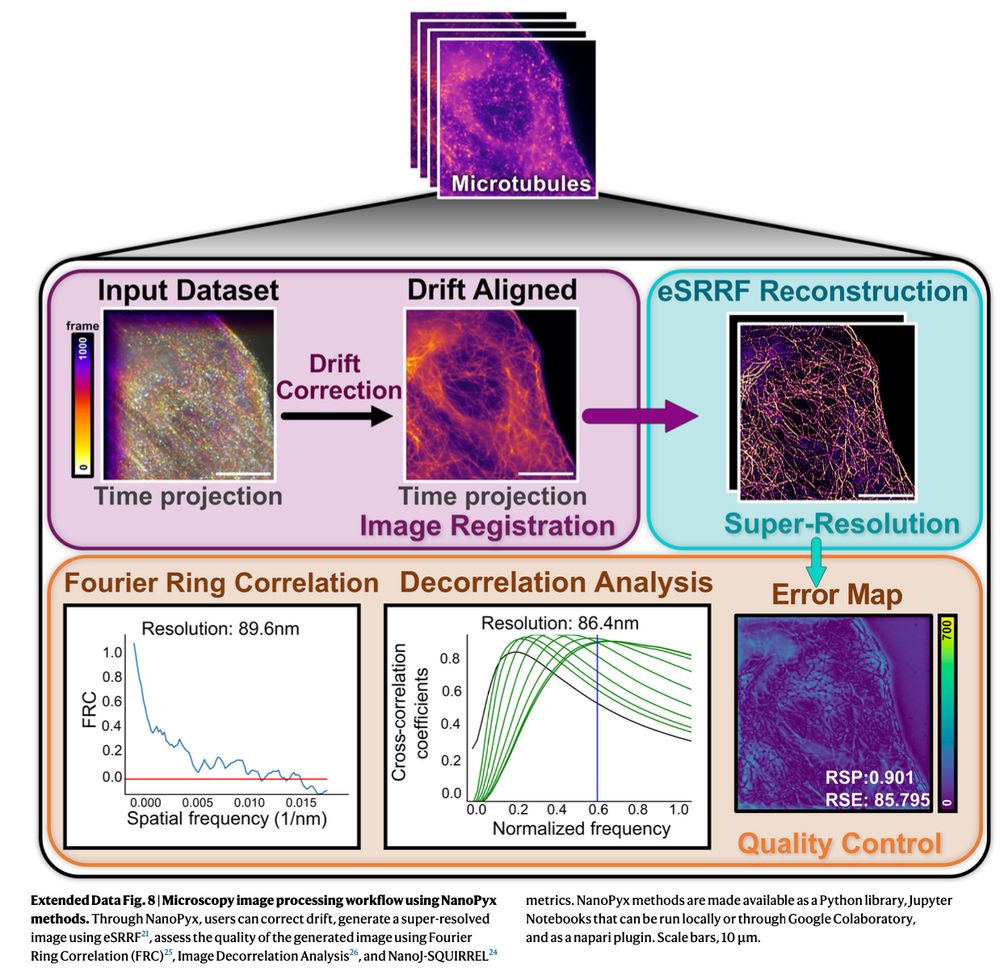
What methods does #NanoPyx offer? We have Image Registration, Denoising, Super-Resolution Image Reconstruction🔬 and Quality Control metrics! (4/n)
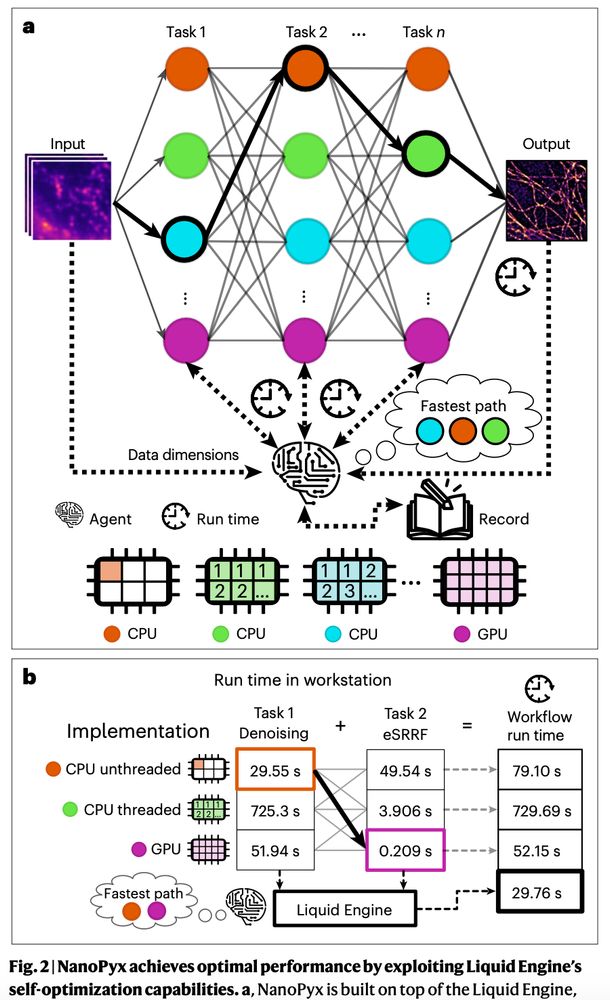
The #LiquidEngine has an AI-powered agent 🤖 that predicts the fastest way of running a specific analysis task, given the input data, the user hardware and task parameters. Incorporating the #LiquidEngine significantly improves the performance of bioimage analysis workflows!(3/n)
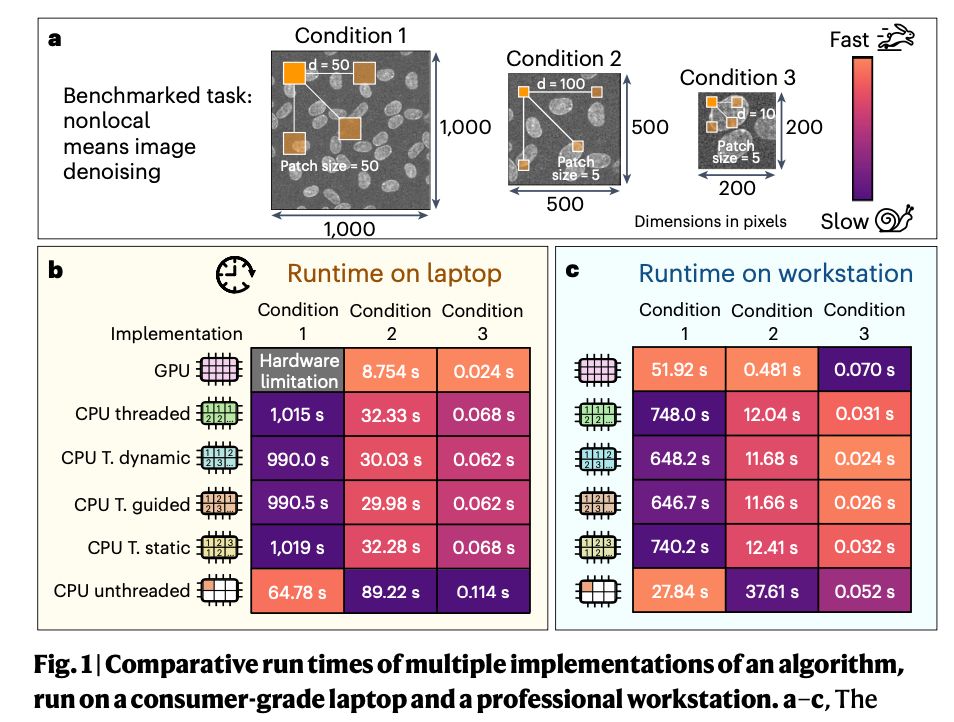
The fastest way of performing any analysis task depends on the user device 💻, the input data 🦠 and the specific task we’re trying to run (2/n).

And to start 2025 strong… Our work #NanoPyx is out in Nature Methods 😍❤️ DOI: doi.org/10.1038/s415...
#NanoPyx is a Python framework that uses the #LiquidEngine to efficiently accelerate bioimage analysis for any user! (1/n) ⬇️