
Now out in @natcomms.nature.com :
versions 2.0 of both BiG-SCAPE and BiG-SLiCE! With significant speed and accuracy increases, as well as new interactive functionalities.
Read the full paper here #openaccess:
www.nature.com/articles/s41...

Now out in @natcomms.nature.com :
versions 2.0 of both BiG-SCAPE and BiG-SLiCE! With significant speed and accuracy increases, as well as new interactive functionalities.
Read the full paper here #openaccess:
www.nature.com/articles/s41...
Aerocavin is an antibiotic with potent and specific anti-Neisserial activity https://www.biorxiv.org/content/10.64898/2026.02.23.707490v1
Unveiling the taxonomic diversity and unprecedented biosynthetic treasure of the phylum Myxococcota https://www.biorxiv.org/content/10.64898/2026.02.12.705523v1
Streptavidins play a multifunctional role within a biotin-pathway antibiotic network encoded in a biosynthetic supercluster https://www.biorxiv.org/content/10.1101/2025.11.10.687665v1

Screening of expanded GRACE mutant collection at six temperatures reveals distinct functional enrichment in genes clustered by fitness profile. Top left: Schematic of GRACE mutant construction. Bottom left: Summary of GRACE mutant collection expansion based on manual curation of biological process gene ontology annotation on Candida Genome Database (CGD). Pie chart with four concentric rings visualizes the relative proportion of genes from various versions of the GRACE collection with biological process descriptions completely unannotated or annotated based on: experimentally characterization, S. cerevisiae ortholog, S. pombe ortholog, orthologs in other fungal species, conserved protein domains, or unannotated; see Materials and methods for full details. Right: Heatmap of genes ordered by hierarchical clustering of essentiality scores across screening temperatures. A score of 4 (red) represents no growth and 0 (yellow) represents growth comparable to wild type.
#Candida albicans is a leading opportunistic fungal #pathogen of humans. Ci Fu, Leah Cowen &co expand the GRACE #Calbicans #FunctionalGenomics resource, identifying genes important for temperature-dependent fitness & highlighting its power to reveal vulnerabilities @plosbiology.org 🧪 plos.io/3Wft0Xg
Very nice Sharok
Thanks Fabrice - you as well!
Congrats Fabrice!
I'm delighted to share a useful FREE web-based app portal of powerful bioinformatics tools for microbial genomes and shotgun microbiome sequences!
It's called Micromics and is built by Middle Author Bioinformatics, led by co-founder @ironark.bsky.social
omix.midauthorbio.com

Comparison of B. theta and E. coli BAM machineries. Only BamAD components are shared, and B. theta BAM has five novel components, BamF-I.
New preprint 🚨: with @madejmar.bsky.social we found that the Bacteroidetes beta-barrel assembly machinery is quite different to E. coli. www.biorxiv.org/content/10.1... #cryo-EM
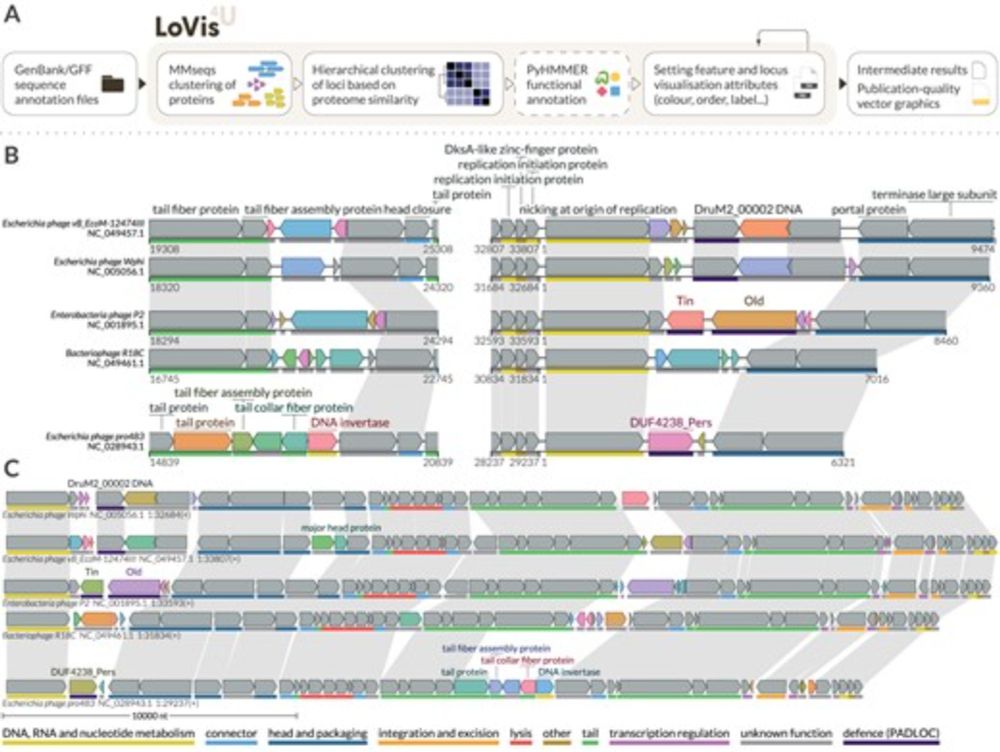
Excited to share that our paper on locus visualisation tool LoVis4u is out! Entirely driven by fantastic PhD student Artyom Egorov @egorov.bsky.social academic.oup.com/nargab/artic...

What do bacterial cells do when they run out of nutrients? Although most bacterial studies focus on cells in exponentially growing states, in the wild bacteria likely spend most of their time slowly starving to death. 1/n
www.biorxiv.org/content/10.1...

Confirmed speakers for the next Great Wall Symposium is coming along! thegreatwall-symposium.org/program/
Abstracts will open next week 15th so start thinking!
September 15-17th, Catania, Sicily
Its a Microbiological opportunity you cant refuse!
A new class of penicillin-binding protein inhibitors to address drug-resistant Neisseria gonorrhoeae https://www.biorxiv.org/content/10.1101/2024.12.27.630553v1
Chemical genomics informs antibiotic and essential gene function in Acinetobacter baumannii https://www.biorxiv.org/content/10.1101/2024.12.05.627103v1
tagging #MicroSky
🚨 PhD opportunity 🚨 The Whelan lab, along w @brockhurstlab.bsky.social, are looking for a PhD student w bioinformatic experience/interest to investigate the evolution of transmissibility in bacterial pathogens, with a focus on P. aeruginosa. More info & link to apply below.
Please spread the word!

Just in time for #WAAW - new collab paper with the Strynadka lab at UBC on structure/function of #peptidoglycan fragment recycling transporter AmpG involved in AmpC-related beta lactam resistance. www.nature.com/articles/s41... @yaegerluke.bsky.social