
www.science.org/doi/10.1126/...
This is clever, very clever
We wrote a perspective "How to build the regulatory genome: a constructionist guide to the cis-regulatory code", out in Development yesterday. Title says it all. Find it here:
journals.biologists.com/dev/article/...
Our most recent work on the “function and evolution” of #nuclear-speckles is now online at Cell @cp-cell.bsky.social
doi.org/10.1016/j.ce...
Read the thread👇 for the highlights of our findings.

Ooooh. Cool new paper on origins of life. A simple 45-nucleotide RNA molecule that can perfectly copy itself.
www.science.org/doi/10.1126/...
Exciting Postdoc Opportunities – Alliance Interinstitutional Program
**Deadline:** March 31, 2026 (5:00 pm CEST)
🔗 Two shared positions: www.syn-gen.de/alliance-pos...
🔗 Full call & application info:: www.health-life-sciences.de/opportunitie...
We are thrilled that our study on the evolution of gene regulation in mammalian cerebellum development – led by @ioansarr.bsky.social, @marisepp.bsky.social and @tyamadat.bsky.social, in collaboration with @steinaerts.bsky.social – is now out in @ScienceMagazine! www.science.org/doi/10.1126/...

“The idea is to put ChatGPT front and center inside software that scientists use to write up their work in much the same way that chatbots are now embedded into popular programming editors.
It’s vibe coding, but for science.”

⚠️ The final work of two former PhD students Till @tschwammle.bsky.social and Verena @verenamutzel.bsky.social is out!
➡️⬅️ They dissect how memory can arise from antisense transcription using mathematical modelling 💻, genomics 🧬 and synthetic biology ⚒️! link.springer.com/article/10.1...
Ah I see, makes sense then! Thanks for the explanation.
I understand the answer to the first question after skimming the paper, but I have to check CREsted first. Though I think my point still stands, it would be more appropriate to compare such a TFBS prediction with something that uses motif+some data, like TF footprinting maybe.
Very interesting work! Is TF-MINDI cell type agnostic? What I mean is does it use the entire S2F model (e.g. all 7000 Borzoi tracks) or when predicting in PMBC you use only the PMBC channels?
Also, I'm not sure if motif enrichment is the right comparison here. S2F models learn from data after all
TF-MINDI is out! A new method to learn cis-regulatory codes through rich embeddings of TF binding sites. TF-MINDI decomposes motif neighbourhoods, and works downstream of any sequence-to-function deep learning model. We deeply study the enhancer code in human neural development, check out the thread

very sad news. Peer Bork was one of the leaders of our field, a wonderful scientist, and he's much too young to be gone. www.embl.org/news/embl-an...
This sounds like it's from a sci-fi novel!

Save the date: April 9 from 4pm to 6pm CET. Our department is hosting an online seminar with @noeliaferruz.bsky.social @sdomcke.bsky.social @const-ae.bsky.social who will talk about models for protein design, large-scale perturbation screens, and benchmarking of perturbation prediction models.

Our paper on the newest version of the Registry of candidate cis-Regulatory Elements (cCREs) is out 🧬
Huge thanks to the many collaborators, experimentalists, analysts and software developers who made this work possible — truly a team effort!
A "meme-torial" of the science is coming soon 👀

The Empire strikes back, not just to grab oil and other riches but, fundamentally, to hide its own weakness at home – and to prepare the ground for subjugating its own people, in Chicago, Portland, NYC etc. Meanwhile, a vassal Europe watches in silence... www.theguardian.com/world/live/2...

📣 I hereby make my Bluesky debut to announce that our work linking DNA binding affinities and kinetics 𝘪𝘯 𝘷𝘪𝘵𝘳𝘰 and 𝘪𝘯 𝘷𝘪𝘷𝘰 for the human transcription factor KLF1 just got published in Cell! @cp-cell.bsky.social
www.cell.com/cell/fulltex...
Key findings in a thread (1/6):
My first @umasschan.bsky.social/@impvienna.bsky.social affiliated paper is up!
tomtom-lite is a re-implementation of tomtom targeting the ML age of genomics. Fast annotations ("what is this motif?") and simple large-scale discovery of motifs.
Check it out!
academic.oup.com/bioinformati...
Hey authors! Check to see if Anthropic stole your book to train their slop generator on. You’re entitled to $1500 per stolen Work.
Look up your work, and if you’re in the database, file a claim
secure.anthropiccopyrightsettlement.com/lookup/

Today I was happy to present Corgi at the @broadinstitute.org Broad Institute at the ML in Drug Discovery Symposium.
If you want to use it for predicting genomic tracks, Corgi is now published on GitHub: github.com/ekinda/corgi

We are excited to welcome @fueyoraquel.bsky.social, @sedonamurphy.bsky.social, and @jmstein.bsky.social as new group leaders at our institute!
Read more about their work:
--> www.molgen.mpg.de/2025-10-31-n...
All three are recruiting in the IMPRS PhD Call!
--> www.molgen.mpg.de/IMPRSPhDproj...

Last chance to turn it off.
On Monday, November 3rd, Microsoft will start using your LinkedIn data for AI training. And remember, you're opted in by default.
To toggle it off 👉 Account - Settings & Privacy > Data privacy > Data for Generative AI Improvement.

The genetic code is full of synonymous codons that, for decades, were assumed to be interchangeable.
Today, with NVIDIA, co-led by @genophoria.bsky.social, we announce CodonFM, a family of open-source AI models that reveal the grammar underlying codon choice: developer.nvidia.com/blog/introdu...

What is a promoter? And how does it work?
We very happy to share our latest work trying to understand enhancer-promoter compatibility.
I am very excited about the results of @blanka-majchrzycka.bsky.social, which changed the way I think about promoters
www.biorxiv.org/content/10.1...
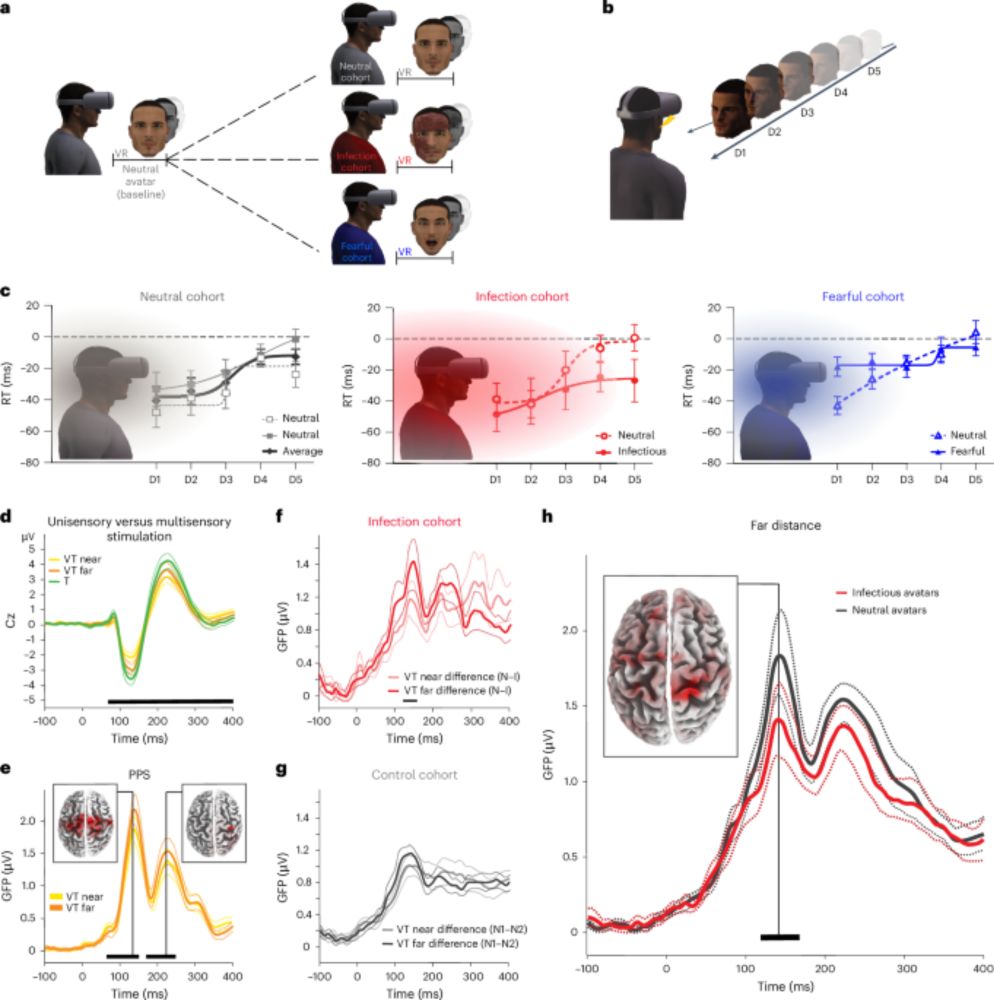
Neural anticipation of virtual infection triggers an immune response
www.nature.com/articles/s41...

⚠️ Paper alert: Using a novel CRISPR screening approach, we mapped the entire regulatory network controlling Xist—key for X-chromosome inactivation.
👉 We discover how sex and development signals are decoded at a single gene locus.
www.nature.com/articles/s41...
👇 Bluetorial

🧵1/ Excited to share our new paper introducing a new #singlecell assay: scTF-seq, a high-throughput single-cell approach to explore how transcription factor (TF) dose shapes cell identity and reprogramming outcomes. 🔗 www.nature.com/articles/s41... Big congrats to the entire team @EPFL & @SIAT_China
⚡⚡Excited to announce I'll be starting my lab at the Max Planck Institute of Molecular Genetics (@molgen.mpg.de) in Berlin in December! Leaving sunny California to join a fantastic environment with colleagues who do super cool work.
🔬🦠I'm hiring at all levels! 🔬🦠Check: www.molgen.mpg.de/fueyo-lab




It's conference week in Cambridge. Yesterday I presented Corgi in the MIT/MGB AI Cures conference, and today I'm at the Broad Institute for the Variant to Function conference.