Congrats!
Congrats!

Here is the @scripps.edu lay audience friendly article that accompanies it.
www.scripps.edu/news-and-eve...
This was a gargantuan but incredibly satisfying collaboration, which was critically supported by a BRAIN Initiative grant, among other funding mechanisms.

Transcriptional and translational dynamics in CA3 vs CA1 neurons. Highlighted in red are ribosomal proteins, and elongation factors
Finally, we found that CA3 neurons translate at higher levels compared to CA1, despite CA1 expressing more transcripts. Many highly translated genes are ribosomal machinery genes, resulting in a different translational landscape that could matter for hippocampal function.

Cadm2 highly-translated isoform has a longer 3'UTR that hosts nELAVL binding sites
Excitingly, we found that, for the same gene, the isoform displaying the highest translation has a longer 3'UTR than the lowest-translated isoform. This seems to result from complex interactions among splicing and negative and positive translational regulators (e.g., microRNAs and nELAVL).


Plasticity genes Grin2b and Celf4 are highly translated in a subset of neurons, but the RNA level is uniform.
We performed simultaneous single-cell transcriptional and translational profiling of the mouse hippocampus. We found that post-transcriptional mechanisms 'clean up' transcriptional noise. We identified a subset of neurons at high translational state, possibly a proxy for plasticity.
Paired with long-read RNAseq, Ribo-STAMP can measure the translation potential of transcript isoforms. Since defects in alternative splicing are strongly linked to neuropsychiatric disorders, understanding how protein production is affected could suggest novel disease mechanisms.

The RNA editor APOBEC1 is fused to a ribosomal protein. During translation, Ribo-STAMP leaves a mark in the form of a C-to-U edit.
Brain cells display the highest levels of post-transcriptional gene expression regulation. Yet, to understand the underlying mechanisms, we need tools that match their incredible cell diversity. The Yeo lab developed Ribo-STAMP to measure translation at single-cell resolution.

Chuffed to see our paper with @geneyeo.bsky.social finally published in @nature.com.
www.nature.com/articles/s41...
Very grateful to Fede, Susie, and Dave from my lab, and Sammi, Eric, and Pratibha from Gene's lab. What did we find? ⬇️ ⬇️ ⬇️
Congrats!
Congrats Alex!
🎉🎉
Come to the Symposium
Congrats!

In our first pitch session, Ryan Shenvi, PhD, @lippilab.bsky.social, PhD, and Li Ye, PhD, will each share their approach to understanding the brain—from neuroactive compound synthesis, to microRNAs in circuit development, to brain energetics—and will take questions after each of their pitches.
Congrats!
Congrats!
Yeah Marko!
Congrats
🎉
First author @norjin.bsky.social has a different version of the birth of this project. A classic case of convergent evolution.
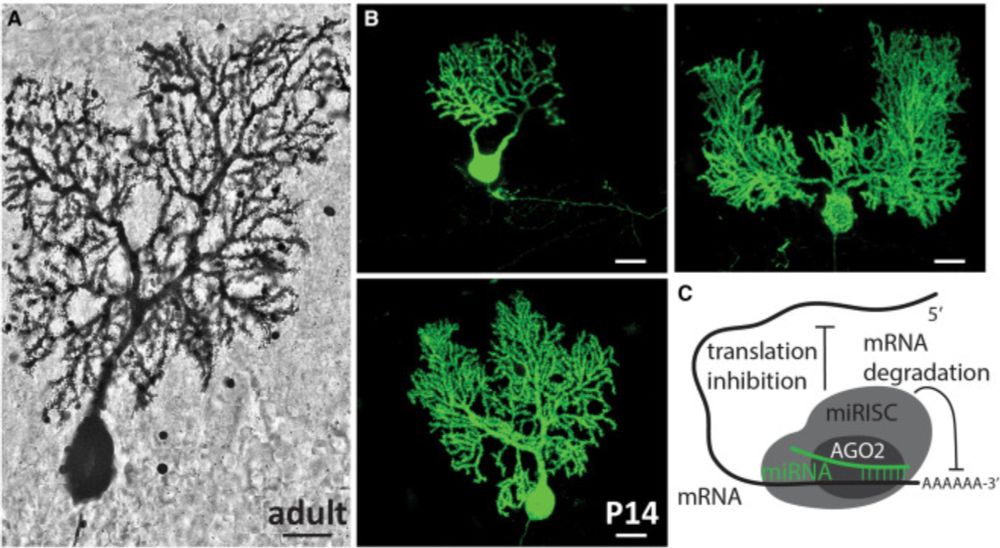
How fitting that the Preview of our Neuron paper was written by the one and only Roy Sillitoe. I saw him giving a talk at the 2018 Neural Development GRC, and his incredible data, storytelling, and enthusiasm convinced me to use Purkinje Cells as a model system.
www.cell.com/neuron/fullt...
Wow! Great stuff Debby congrats!
😂
Congrats buddy! 🎉🎉
Congrats!
Congrats Alex!
Congrats Chen!
Congrats Emilia!

Postdoc opportunity in the heart of Europe! Check out the flyer below.
Join our team on an international project with Anthony Holtmaat exploring synaptic plasticity and specificity in cortical and thalamocortical circuits.