Highly appreciate your valuable comments and suggestions @kamounlab.bsky.social @thorstenlangner.bsky.social @joewin
Highly appreciate your valuable comments and suggestions @kamounlab.bsky.social @thorstenlangner.bsky.social @joewin
Using unused RNA-seq data from wheat blast–infected field samples (via Open Wheat Blast), we identified candidate S genes and validated one by in-planta experiments. Thanks @tofazzalislam for collaboration.
Special thanks @kamounlab.bsky.social for leading OpenWheatBlast. www.nature.com/articles
You’re most welcome. You were the one who generously taught me GE through one-on-one sessions, and I’ll always be grateful for that. 🙏
Congratulations Robert ❤️🙏
Definitely worth a read.
Big thanks to @KamounLab for highlighting key messages! 🙌
Helped us see more clearly how science should be done—beyond journals and impact factors.
Thanks again @Edel_PLopez for your insights 🙏.


For the first time in my 🇩🇪 life Deutsche Bahn was on time today🫡
Hello, beautiful Köln!
#ISMPMI2025
Mid-week #mycorrhiza 🍄
View through the 3D structure of an arbuscule. This is in a rice root cell colonised by the arbuscular mycorrhizal fungus Rhizophagus irregularis 🔬
Thanks for your exciting insights about science and communication during ML 🙏👍
Here is our latest study @newphyt.bsky.social , led by @fabianvanbeveren.bsky.social and Yvet Boele! Our first dive into ectomycorrhizae!
nph.onlinelibrary.wiley.com/doi/10.1111/...
Do you want to learn about the convergent evolution of ECM? Have a look at the thread prepared by Fabian🔽
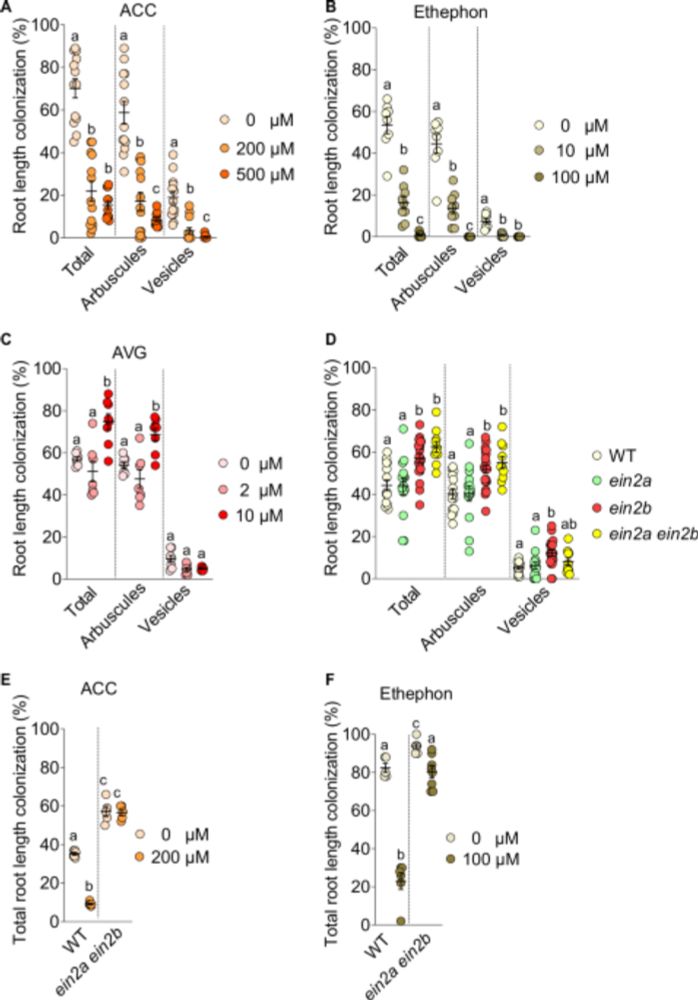
New #PlantSci paper by team around @carogutj.bsky.social @mpi-mp-potsdam.bsky.social in @naturecomms.bsky.social
doi.org/10.1038/s414...