
SPIDR enables multiplexed mapping of RNA-protein interactions and uncovers a mechanism for selective translational suppression upon cell stress @cellcellpress.bsky.social @mitchguttman.bsky.social
www.cell.com/cell/fulltex...

SPIDR enables multiplexed mapping of RNA-protein interactions and uncovers a mechanism for selective translational suppression upon cell stress @cellcellpress.bsky.social @mitchguttman.bsky.social
www.cell.com/cell/fulltex...
This work was led by co-first authors Jimmy Guo and Erica Wolin with support from amazing teams from our lab @caltech.edu and the Jovanovic lab @columbiauniversity.bsky.social and @jbquerido.bsky.social lab with financial support from NHGRI, NSF, @genometdcc.bsky.social.

We used SPIDR to identify mTOR-dependent changes and observed that 4EBP1 showed a dramatic increase in binding upon mTOR inhibition specifically at mRNAs containing a TOP-motif, suggesting a new model for how translational repression is selectively achieved upon mTOR inhibition.

We identified an interaction between LARP1 and 18S rRNA located within the mRNA channel entry site on the 40S small ribosomal subunit and @jbquerido.bsky.social resolved this structure at 2.8 Å using single-particle cryo-EM.

Single nucleotide binding maps generated by SPIDR can map known RNP structures at atomic resolution and identify novel components within RNP structures.

We show that SPIDR generates high quality data across a diverse range of RBPs, including transcription, splicing, translation, and miRNA biogenesis, all within a single experiment.

SPIDR uses a dramatically simplified split-and-pool based strategy to increase the throughput of CLIP by two orders of magnitude. SPIDR enables the rapid generation of consortium-level datasets within any molecular biology lab without the need for specialized training or equipment.

Many proteins bind RNA, yet we still don’t know what RNAs most bind because methods map one RBP at a time. In @cp-cell.bsky.social, with the Jovanovic lab, we describe SPIDR – a method for mapping the RNA binding sites of dozens of RBPs in a single experiment. www.sciencedirect.com/science/arti...
This work was led by co-first authors Drew Perez and Isabel Goronzy and our amazing team @caltech.edu, and funding from NHGRI.

By enabling the generation of consortium-level datasets within any molecular biology lab, ChIP-DIP facilitates a fundamental transition from ‘reference-maps’ to context-specific maps and represents a transformative new tool for dissecting cell-type specific gene regulation.

We used ChIP-DIP to explore quantitative combinations of histone modifications that define distinct classes of regulatory elements and integrated these signatures with regulatory factor binding to identify their functional activity.

We used ChIP-DIP to measure temporal chromatin dynamics in primary mouse dendritic cells following stimulation and correlate these with transcriptional changes. For this, we mapped multiple time points within a single experiment, multiplexing across both proteins and samples.

ChIP DIP generates highly accurate maps for ALL classes of DNA-associated proteins, including histone modifications, chromatin regulators, transcription factors, and RNA polymerases.

ChIP-DIP uses a simple antibody-labeling strategy followed by split-and-pool barcoding to multiplex DNA-protein mapping. This enables the rapid generation of consortium-level datasets within any molecular biology lab without the need for specialized training or equipment.
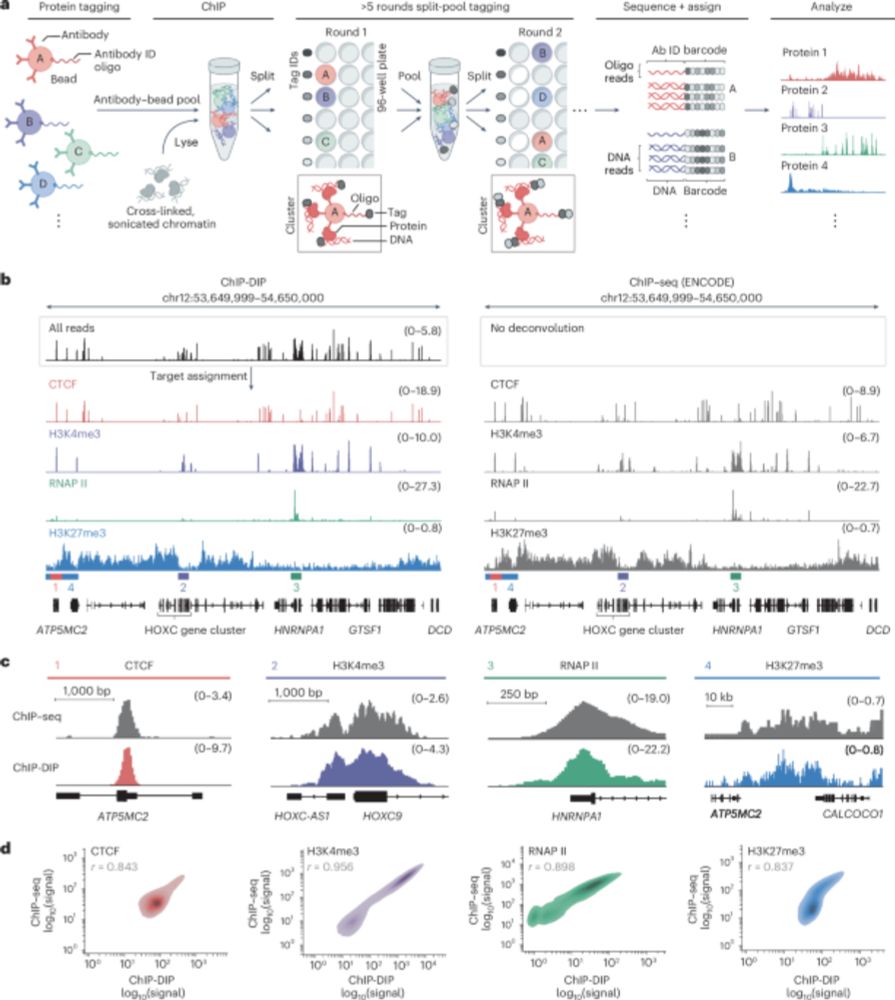
Gene regulation involves thousands of proteins that bind DNA, yet comprehensively mapping these is challenging. Our paper in Nature Genetics describes ChIP-DIP, a method for genome-wide mapping of hundreds of DNA-protein interactions in a single experiment.
www.nature.com/articles/s41...
Great thread on our paper from Jan👇

A report made counterintuitive claims that PRC2 binds
→ more RNA than PTBP1 & hnRNPU
→ RNAs that do not exist in the cell
We found a simple explanation: the authors ignore most RNA in the sample, leading to inflated background & misleading conclusions.
www.biorxiv.org/content/10.1...
Leonid Mirny and I wrote this for all interested in chromosomes: "The chromosome folding problem and how cells solve it"
www.cell.com/action/showP...