This is beautiful.
This is beautiful.
i'm just attending, will try book earlier next year then :)
is there still available ticket by any chance ? lol

Imagine a brain decoding algorithm that could generalize across different subjects and tasks. Today, we’re one step closer to achieving that vision.
Introducing the flagship paper of our brain decoding program: www.biorxiv.org/content/10.1...
#neuroAI #compneuro @utoronto.ca @uhn.ca
Big congrats !
Very happy to see that Pleias multilingual data processing pipelines have contributed to the largest open pretraining project in Europe.
From their tech report: huggingface.co/swiss-ai/Ape...

Not that it comes as much of a surprise to many of us, but it's worth emphasizing once again - the 👏 brain 👏 uses 👏 distributed 👏 coding 👏. 😁
Two new papers from the #IBL looking at brain-wide activity:
www.nature.com/articles/s41...
www.nature.com/articles/s41...
#neuroscience 🧪
Stunning cryo-ET from Peijun Zhang lab: Direct visualization of HIV-1 nuclear import!
Hundreds of viral cores captured entering the nucleus. The NPC dilates to let the capsid through. A masterclass in correlative microscopy that makes it quantitative. A leap for structural virology! @emboreports.org

Internship Position on the Lattice Estimator martinralbrecht.wordpress.com/2025/08/27/i...
In short, world-class research from NeurIPS accessible in Europe.
EurIPS takes place over 3 days + 2 workshop days, at the same time as NeurIPS in San Diego.
Follow this account for more updates, and see you in wonderful Copenhagen 📅
eurips.cc

Emmanuel Levy @elevylab.bsky.social has joined BlueSky 🌟 with a fantastic Cell paper with Shu-ou Shan, showing the interactome of the TOM complex and how cotranslational mito import prioritizes large globular domains. Beautiful science!
Give him a warm welcome 🎉
link: www.cell.com/cell/fulltex...

EurIPS includes a call for both Workshops and Affinity Workshops!
We look forward to making #EurIPS a diverse and inclusive event with you.
The submission deadlines are August 22nd, AoE.
More information at:
eurips.cc/call-for-wor...
eurips.cc/call-for-aff...
Excited to be at ACL! Join us at the Table Representation Learning workshop tomorrow in room 2.15 to talk about tables and AI.
We also present a paper showing the sensitivity of LLMs in tabular reasoning to e.g. missing vals and duplicates, by @cowolff.bsky.social at 16:50: arxiv.org/abs/2505.07453

Bootstrapped a Private GenAI Startup to $1M revenue, AMA
blog.helix.ml/p/bootstrapp...
![What do representations tell us about a system? Image of a mouse with a scope showing a vector of activity patterns, and a neural network with a vector of unit activity patterns
Common analyses of neural representations: Encoding models (relating activity to task features) drawing of an arrow from a trace saying [on_____on____] to a neuron and spike train. Comparing models via neural predictivity: comparing two neural networks by their R^2 to mouse brain activity. RSA: assessing brain-brain or model-brain correspondence using representational dissimilarity matrices](https://cdn.bsky.app/img/feed_thumbnail/plain/did:plc:e6ewzleebkdi2y2bxhjxoknt/bafkreiav2io2ska33o4kizf57co5bboqyyfdpnozo2gxsicrfr5l7qzjcq)
What do representations tell us about a system? Image of a mouse with a scope showing a vector of activity patterns, and a neural network with a vector of unit activity patterns Common analyses of neural representations: Encoding models (relating activity to task features) drawing of an arrow from a trace saying [on_____on____] to a neuron and spike train. Comparing models via neural predictivity: comparing two neural networks by their R^2 to mouse brain activity. RSA: assessing brain-brain or model-brain correspondence using representational dissimilarity matrices
In neuroscience, we often try to understand systems by analyzing their representations — using tools like regression or RSA. But are these analyses biased towards discovering a subset of what a system represents? If you're interested in this question, check out our new commentary! Thread:

It was great to work on the ODAC25 paper with the Meta FAIR Chemistry and Georgia Tech. A leap forwards in modelling direct air carbon capture with metal organic frameworks, with much better data and larger models.
Paper: arxiv.org/abs/2508.03162
Data and models: huggingface.co/facebook/ODA...

1/ Excited to share a new preprint!
Our latest study uncovers how serotonin precisely controls the “time window” for fear learning, ensuring that our brains link cues (CS) & threats (US) only when it’s adaptive.
#Neuroscience #FearLearning
www.biorxiv.org/content/10.1...
is there a recording on YouTube?

Abstract. The argument size of succinct non-interactive arguments (SNARG) is a crucial metric to minimize, especially when the SNARG is deployed within a bandwidth constrained environment. We present a non-recursive proof compression technique to reduce the size of hash-based succinct arguments. The technique is black-box in the underlying succinct arguments, requires no trusted setup, can be instantiated from standard assumptions (and even when P = NP!) and is concretely efficient. We implement and extensively benchmark our method on a number of concretely deployed succinct arguments, achieving compression across the board to as much as 60% of the original proof size. We further detail non-black-box analogues of our methods to further reduce the argument size.
zip: Reducing Proof Sizes for Hash-Based SNARGs (Giacomo Fenzi, Yuwen Zhang) ia.cr/2025/1446
Very excited to release a new blog post that formalizes what it means for data to be compositional, and shows how compositionality can exist at multiple scales. Early days, but I think there may be significant implications for AI. Check it out! ericelmoznino.github.io/blog/2025/08...
Me too !

Attractors are usually not mechanisms - new blog post: open.substack.com/pub/kording/...

Together with @repromancer.bsky.social, I have been musing for a while that the exponentiated gradient algorithm we've advocated for comp neuro would work well with low-precision ANNs.
This group got it working!
arxiv.org/abs/2506.17768
May be a great way to reduce AI energy use!!!
#MLSky 🧪


👨🎓🧾✨#icml2025 Paper: TabICL, A Tabular Foundation Model for In-Context Learning on Large Data
With Jingang Qu, @dholzmueller.bsky.social, and Marine Le Morvan
TL;DR: a well-designed architecture and pretraining gives best tabular learner, and more scalable
On top, it's 100% open source
1/9

(1/3) Excited to introduce our new GRAB sensors for a series of steroid hormones! These tools enable real-time detection of steroid hormone dynamics in vivo🐭🧠. Happy to share these sensors and welcome any feedback! Please contact yulonglilab2018@gmail.com for information.

1/3) This may be a very important paper, it suggests that there are no prediction error encoding neurons in sensory areas of cortex:
www.biorxiv.org/content/10.1...
I personally am a big fan of the idea that cortical regions (allo and neo) are doing sequence prediction.
But...
🧠📈 🧪

I am very proud to present you the first paper from the main research line of my group at the @mpiage.bsky.social in collaboration with my current group @molepi.bsky.social @bds-lumc.bsky.social at the LUMC, which is now published in @geroscience.bsky.social!
link.springer.com/article/10.1...

Super excited to see this paper from Armin Lak & colleagues out! (I've seen @saxelab.bsky.social present it before.)
www.cell.com/cell/fulltex...
tl;dr: The learning trajectories that individual mice take correspond to different saddle points in a deep net's loss landscape.
🧠📈 🧪 #NeuroAI
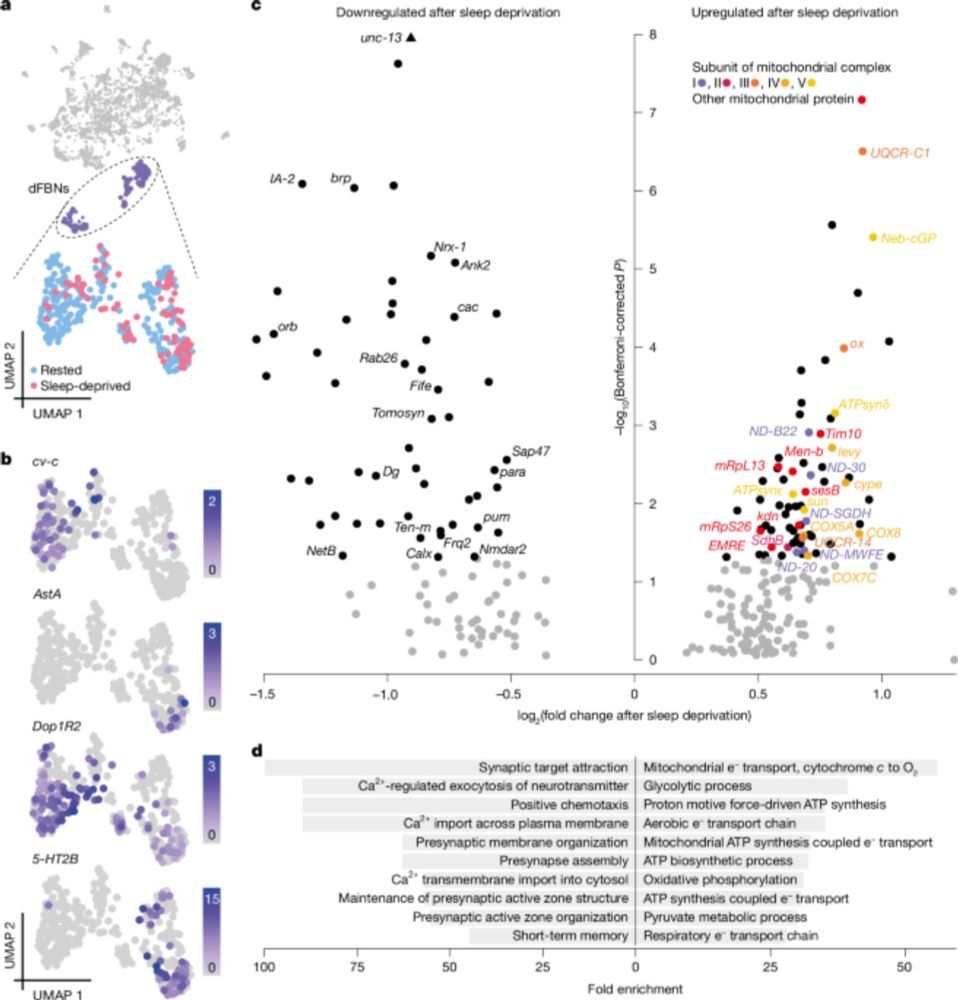
Mitochondrial origins of the pressure to sleep
www.nature.com/articles/s41...

The 1st major study from @arc-cogitate.bsky.social
is out today in @nature.com—a landmark collaboration testing theories of consciousness through rigorous, preregistered science. Data & tools shared openly.
Contributed from @mcgillumedia.bsky.social @theneuro.bsky.social
nature.com/articles/s41...