Find out more: flip.protein.properties
I'm especially happy about continuing to work with an amazing group of scientists. Thanks @kevinkaichuang.bsky.social , @kdidi.bsky.social , Bruce Wittmann, @kadinaj.bsky.social, Maya Czeneszew, @sarahalamdari.bsky.social, @alexijie.bsky.social, @thisismadani.bsky.social, ++
I love this gritty work here; there are no new architectures, no leaderboard-topping number to screenshot. However, it's how we (and hopefully the community) can measure whether the models we're all building and using are getting better where it counts — at the bench.
That's not a criticism to any method. New benchmarks are precisely needed to see where we are at, and set some target of where we could go from here.
Especially on the wild-type and position splits, current transfer learning doesn't consistently win. No single pLM architecture dominates. Scaling hasn't closed the gap yet.
The answer in 2026 is largely the same. Simple ridge regression on one-hot sequences, optionally supplemented with zero-shot pLM likelihoods, often matches or outperforms fine-tuned protein language models.
FLIP2, adds seven new sequence-fitness landscapes - industrial enzymes, nucleases, rhodopsins, protein-protein interactions - and 16 splits that test the generalization axes protein engineers really hit: more mutations, new positions, higher fitness, different wild-types.
We were interested in how things had changed 5 years after our first release. So, we built FLIP2 on select datasets from great labs across the world, many of which have gracefully agreed to make their data freely available.
FLIP spawned fast development of several different benchmarking efforts across protein design, engineering, and variant effect assesment.
The answer in 2021 was: sometimes, but simpler models hold up surprisingly well.
Five years ago, we released FLIP. The core question was: can ML models for protein fitness prediction generalize in the ways that actually matter for protein engineering, i.e. low data, extrapolation to more mutations, out-of-distribution sequences?

We made FLIP2, a protein fitness benchmark spanning seven new datasets, including enzymes, protein-protein interactions, and light-sensitive proteins, as well as splits that measure generalization relevant to real-world protein engineering campaigns.
You can use the model right now to freely generate families for single sequence inputs (i.e., diversification conditioned by intrinsic representations of evolution), or to engineer proteins based on family promts (diversification by conditioning on particular evoluationary trajectories).
In essence, we probed the model's ability to ricapitulate family statistics, bootstrap protein structure prediction, and assess mutation effect, demonstrating excellent performance across all tasks, especially using test-time-scaling via prompt conditioning.
With ProFam-1, we scaled learning from single sequence to protein family definitions of different kinds, curating a large protein family corpus, ProtFam-atlas. I'm particularily stoked about the idea of inference-time-compute. This contribution laid out a very exciting path for future work.
Our latest protein family-based GenAI collection of tools and datasets, ProFam, is out now. Everything -- from data, training and inference code, to a 215M llama-based ProFam-1 are fully open sourced.
🧵

Another exciting opportunity, this time as a colleague at Duke! Join as tenure track assistant prof. in Cell Bio & let’s work on closing the gap between in-silico and in-vivo: www.nature.com/naturecareer...
Important: application closes Nov 1st!!!

Another opening: Senior Multiscale Biology Applied Research Scientist!
nvidia.eightfold.ai/careers/job/...
Are fascinated by fundamental data modalities across biology like RNA-seq, mass spec & want to build computational tools that harnessing data to build intelligence?
Come: join the team!
I don’t dare question the HR gods about their designs :)

Are you passionate about leading collaborative, fast moving, applied bioinformatics research projects that help the entire community move forward?
Apply to work in my team at NVIDIA: nvidia.eightfold.ai/careers/job/...
I should add: structure prediction inference is INCREDIBLY efficient for the form factor and power profile.
It's an inference monster.
Structure prediction on it works.
Design to come.
This will be updated later today... research.nvidia.com/labs/dbr/ass...

Great talk by @machine.learning.bio at the 5th Virtual @chembiotalks.bsky.social. He talked about the use of #MachineLearning and #BigData to address biological questions. Cool insights into both predicting functions and designing proteins
ieeexplore.ieee.org/document/947...
arxiv.org/abs/2503.00710
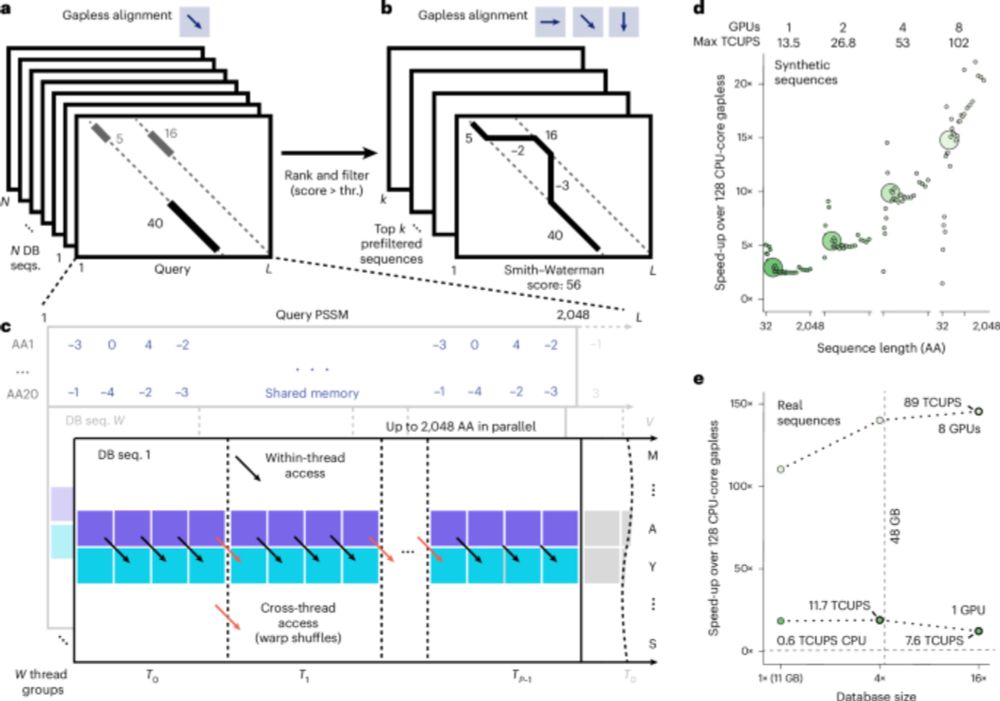
GPU-accelerated MMseqs2 offers tremendous speedup for homology retrieval, protein structure prediction with ColabFold, and protein structure search with Foldseek. @martinsteinegger.bsky.social @milot.bsky.social @machine.learning.bio
www.nature.com/articles/s41...
Looking forward to hearing about the potential of machine learning for #Biology and #DrugDiscovery from an industry perspective. Register for the Virtual @chembiotalks.bsky.social to hear the perspective of Chris Dallago (@machine.learning.bio) from Nvidia.
#ChemBio #Chemsky #ML #MachineLearning
I still feel criminal for the handle but the manuscript embodies it well.
Moore’s law applied to speed not accuracy. I don’t think fundamentally the discoveries we are after are entirely dependent on speed.
I think the better law here is garbage in garbage out.
In that sense, you can wait for better data/curation, but it’s also fun to take destiny in your own hands :)

(1/5) Venoms are a vast, largely untapped library of bioactive molecules—and our new paper in @natcomms.nature.com @natprot.nature.com reveals just how powerful they can be. 🐍⚡️

Excited to have participated in the 2025 Symposium on Generative AI in Molecule Discovery in beautiful Munich, along with amazing scientists and colleagues @machine.learning.bio, Francesca Grisoni, @ewaszczurek.bsky.social, @fabiantheis.bsky.social and more... 🔬🤖 events.hifis.net/event/2015/
Folddisco finds similar (dis)continuous 3D motifs in large protein structure databases. Its efficient index enables fast uncharacterized active site annotation, protein conformational state analysis and PPI interface comparison. 1/9🧶🧬
📄 www.biorxiv.org/content/10.1...
🌐 search.foldseek.com/folddisco