New preprint by @traver.bsky.social . Congrats Rosalind, Traver and colleagues!
New preprint by @traver.bsky.social . Congrats Rosalind, Traver and colleagues!
and our extensive exploration of scoring isogenic CRISPR screens @donnellycentre.bsky.social @sickkidsto.bsky.social www.biorxiv.org/content/10.1...
This chemical-genetic interaction map complements our efforts to map genetic interactions: www.biorxiv.org/content/10.1...
And more isogenic CRISPR screens coming out of Toronto @sickkidsto.bsky.social, this time by Mike Tyers and team - congrats! www.biorxiv.org/content/10.6...
Great new work from @colmr.bsky.social predicts a cancer dependency map for paralogs: doi.org/10.64898/202....
Perhaps one of the most anticipated data in my recent memory- amazing talk by Carmen from @sangerinstitute.bsky.social at @eurocancerdepmap.bsky.social
Truly inspiring talk by @icrlordlab.bsky.social at @eurocancerdepmap.bsky.social

TF-MAPS: fast high-resolution functional and allosteric mapping of DNA-binding proteins by @XianghuaLi2
Are Transcription Factors really 'undruggable'?
www.biorxiv.org/content/10.1...
Thank you @ioriolab.bsky.social for having me. It was great to talk science. @humantechnopole.bsky.social is truly a magnificent place to do research.
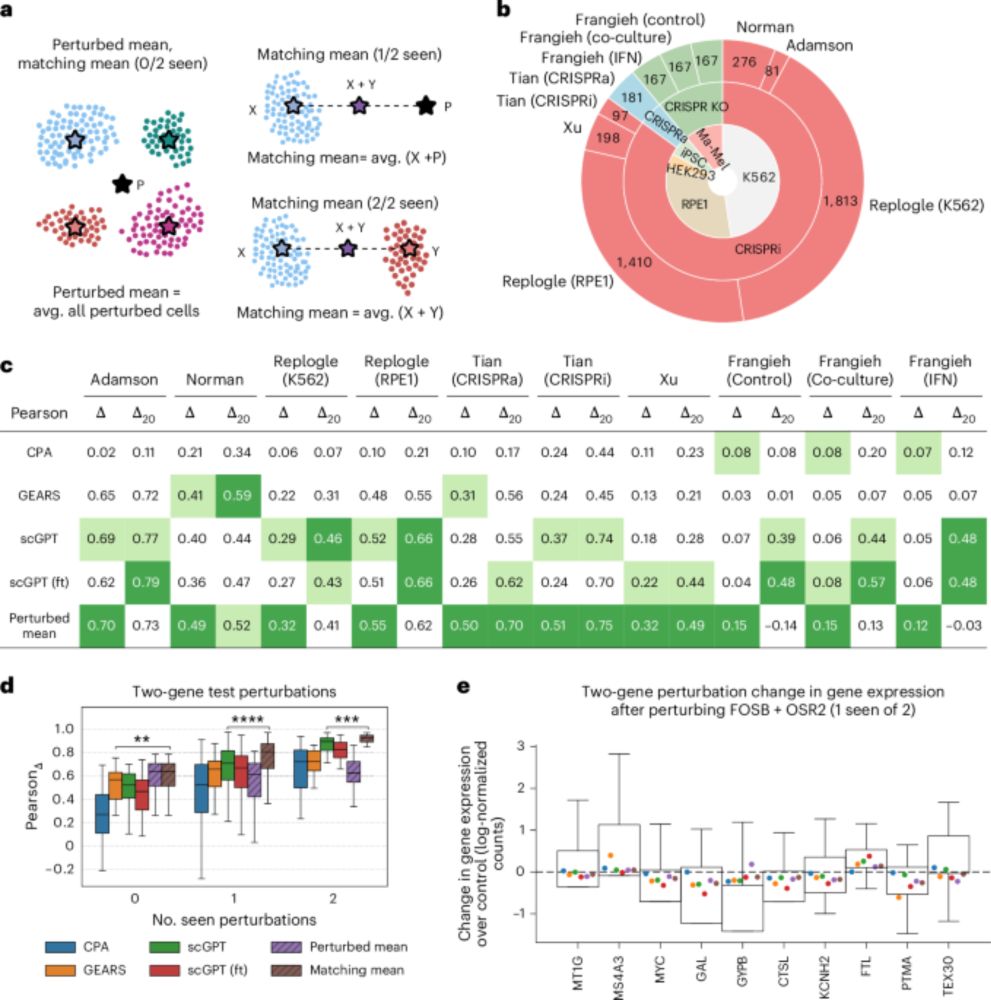
"...predicting responses to unseen perturbations is substantially harder than standard metrics suggest". I am not surprised. www.nature.com/articles/s41...

Can you find therapeutic targets for cardiomyopathies using simple, clean genetics in a human cell line? www.nature.com/articles/s41...

Online now: Combined MEK and PARP inhibition enhances radiation response in rectal cancer
Thank you @colmr.bsky.social for the shoutout! It was quite a journey. If it is only nearly as useful as the @depmap.org we are happy. In any case, it complements this data ...as expected!?

I’m excited to highlight our latest paper, just published in Cell 🎉
We report the existence of a previously uncharacterized signaling pathway that is responsible for activating cell death upon loss of gene expression.
1/n 🧪
www.cell.com/cell/fulltex...

Variant characterization in the intrinsically disordered human proteome - here is how we did it using short linear motif prediction, AlphaFold, and experimentation. Check out our preprint: www.biorxiv.org/cgi/content/...
One thing that really bothers me with the new "virtual cell" terminology is that it is currently largely focused on a very narrow definition of models that can predict effects of trans perturbations (gene dosage, drugs etc) on gene expression. 1/
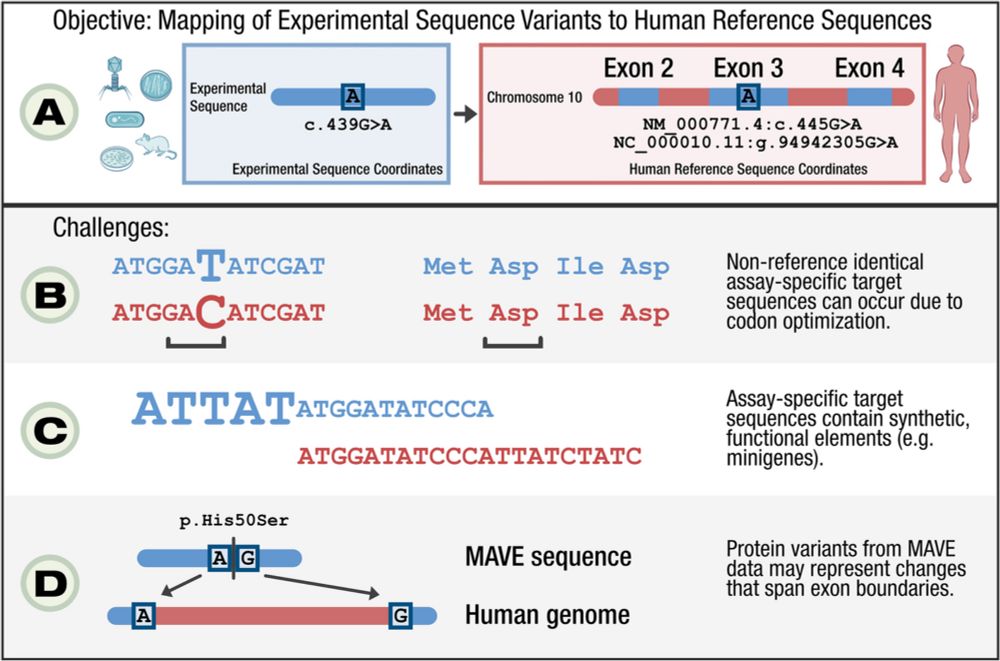
We are pleased to announce the publication of our manuscript "Mapping MAVE data for use in human genomics applications" in Genome Biology! 🧵 1/3 genomebiology.biomedcentral.com/articles/10....
agree.
Our "Atlas of Variant Effects 2030 Roadmap" is live: zenodo.org/records/1542...
1/n

Jason Moffat at #VariantEffect25

Have you discovered the Night Science Podcast yet?
@nightsciencepod.bsky.social
Kind words from one of the up-and-coming leaders of the field ;)
Thanks @colmr.bsky.social ! I felt this was a suitable place to recognize the substantial efforts that go into perturbation experiments.
I appreciate the recognition! I guess pertomics mostly describes a reverse genetics view and "buffomics" would place the phenotype in the center ;)