
💻 GitHub: github.com/anton-bushui...
📄 Paper: arxiv.org/abs/2411.02109

💻 GitHub: github.com/anton-bushui...
📄 Paper: arxiv.org/abs/2411.02109

I am super excited that our paper on inter-laboratory comparability of non-targeted LC–MS/MS analysis of dissolved organic matter was just published in ES&T.
pubs.acs.org/doi/10.1021/...

Are you planning to attend the @appliedmldays.bsky.social conference at EPFL? Together with @arianemora.bsky.social we are organizing an ML4Enzymes workshop on Feb 12 morning, including expert presentations and a hands-on coding session. Full program: docs.google.com/document/d/1...

🧬 A new paper from the group of Ed Curtis at IOCB Prague in collab with the @pluskal-lab.org introduces a structure-guided way to explore nucleic acid sequence space that goes beyond traditional random mutagenesis.
#research #machinelearning #nucleicacids #chemistry
1/3

📰 @forbescesko.bsky.social ve svém výběru českých vědeckých objevů zmínil také výzkum z našeho ústavu.
Vedle dalších projektů upozornil na práce týmu Petra Cíglera a týmu @pluskal-lab.org ► forbes.cz/tricko-proti... 📲
#ceskaveda

Excited to release BoltzGen which brings SOTA folding performance to binder design! The best part of this project is collaborating with a broad network of leading wetlabs that test BoltzGen at an unprecedented scale, showing success on many novel targets and pushing the model to its limits!
We train machine learning models on millions of proteins. But when it comes to making predictions, do we need them to understand all proteins at once? Often, we need an accurate model for the specific protein we are studying or designing. We address this with ProteinTTT arxiv.org/abs/2411.02109 1/🧵
BIG BIG congratulations to our PhD student
@roman-bushuiev.bsky.social for receiving the Google PhD Fellowship 2025 in Health Research! 🎉💰 goo.gle/43wJWw8

Je život ve vesmíru vzácný, nebo se naopak může vyskytovat skoro všude?
Odpověď na to hledá projekt #PROTOCELL Kláry Hlouchové a @pluskal-lab.org, který vzniká v @iocbprague.bsky.social. 1/3
#ČeskáCestaDoVesmíru #CzechSpace #CzechSpaceWeek2025 #CzechSpaceWeek #UOCHB #IOCB #IOCBPrague
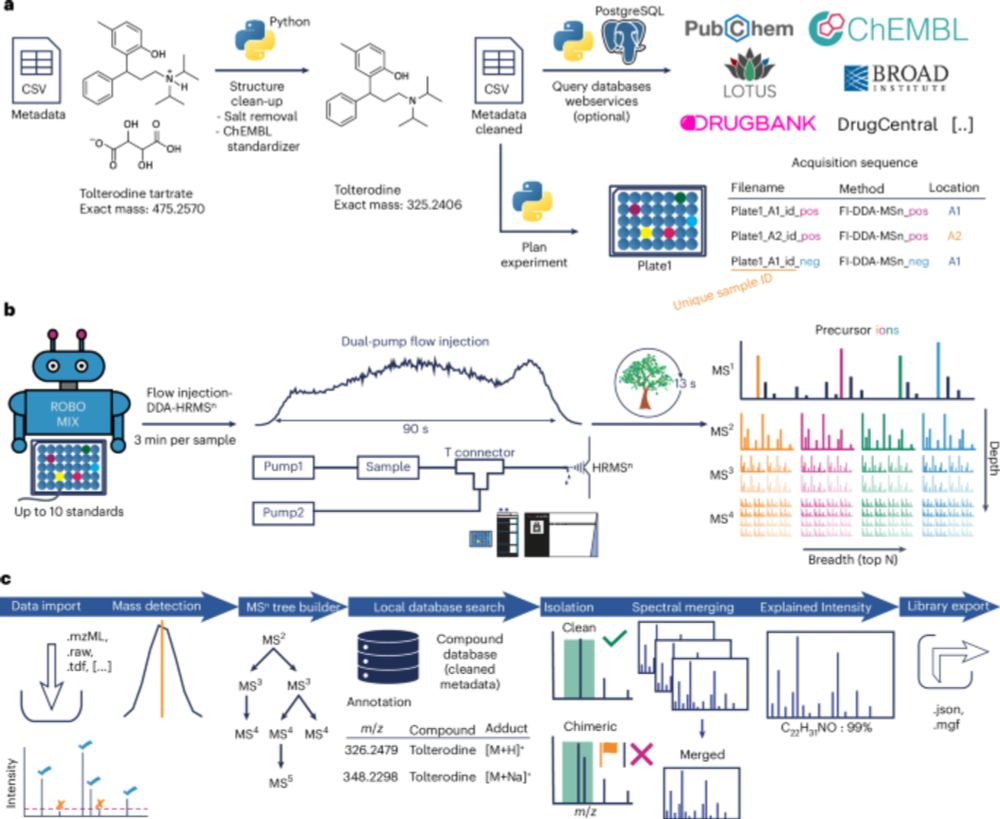
MSnLib: a large-scale, open multi-stage fragmentation mass spectral library for 30,000 unique small molecules.
@robinschmid.bsky.social @pluskal-lab.org
www.nature.com/articles/s41...
Also, it is a great showcase of our ongoing collaboration with @mzio-gmbh.bsky.social @robinschmid.bsky.social on developing new @mzmine.bsky.social workflows.
Most importantly, the effort still continues and we hope to reach 200,000 analyzed compounds in a few months, with all the data publicly available with no restrictions. This open resource is super valuable for the whole metabolomics community and especially for ML applications.
I am very proud of the final outcome of this project, mainly driven by Corinna Brungs and supported by numerous labs who donated compounds for analysis.




The first day of our bioML symposium at @iocbprague.bsky.social went very well. It is amazing to see how fast this field is moving forward. Thanks to everyone for the fantastic talks! 🤖🧬
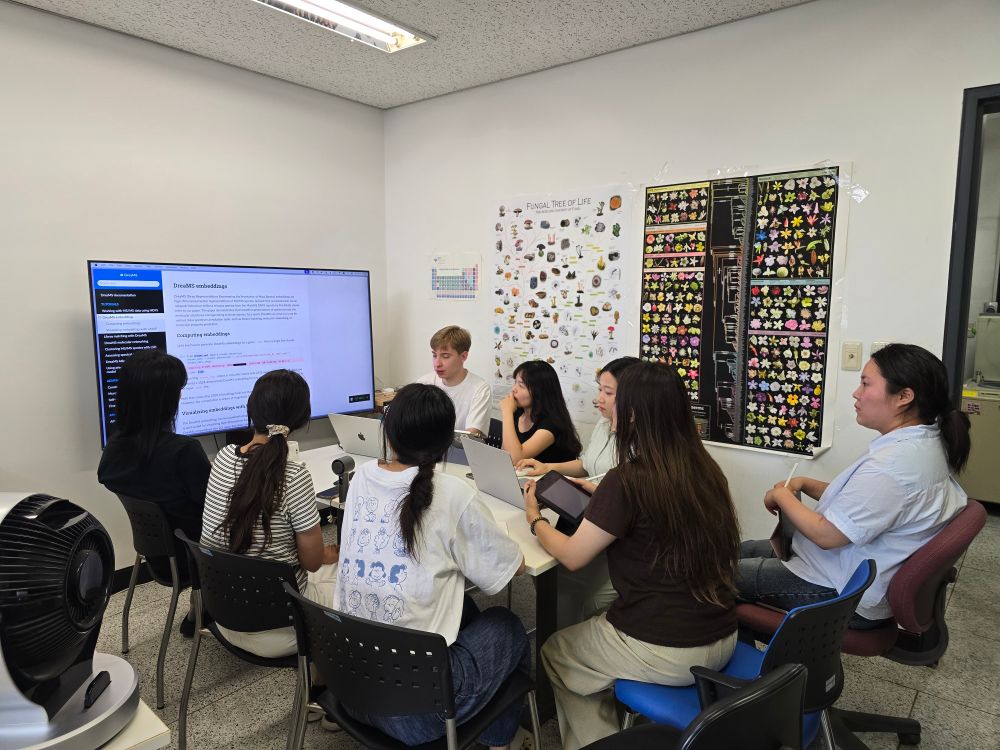

Yesterday @roman-bushuiev.bsky.social visited our lab. He gave a seminar talk about his recent work on DreaMS (www.nature.com/articles/s41...), and provided a hands-on training to my lab members. Looking forward to what we will discover using this awesome tool!
My pleasure! 🙂
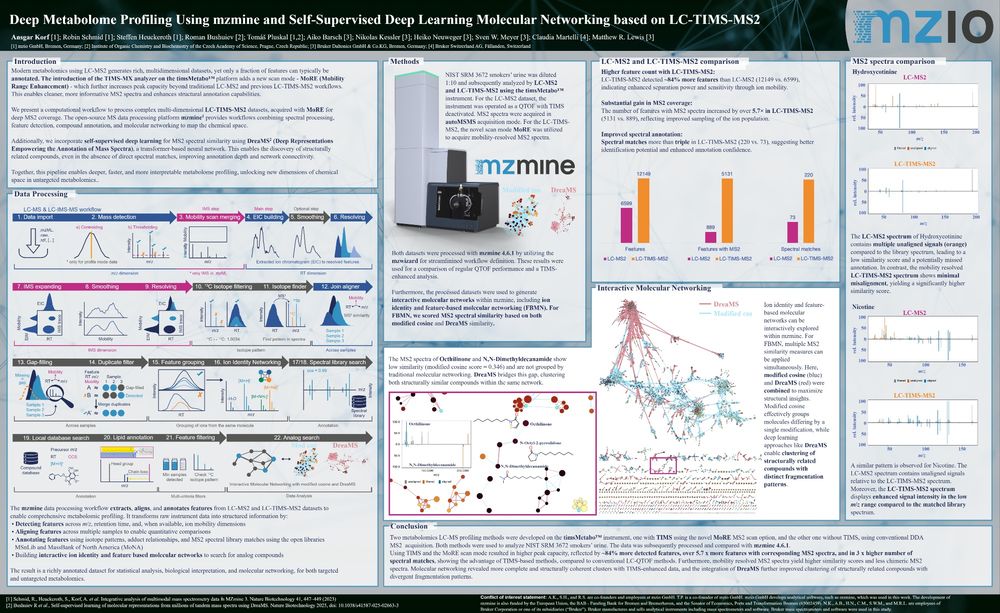
At #ASMS2025, we presented a next-gen workflow pairing the #timsMetabo MoRE (Mobility Range Enhancement) with #mzmine and #DreaMS
Compared to conventional LC-MS2, the data delivered:
- 84% more detected features
- 5.7× more MS² spectra
- 3× more spectral matches
Read the full poster to learn MoRE
🚨 MetaboART 🎨 2025 – Deadline Extended:
🗓️ Submissions now open until June 13!
Two categoried:
👤 Human-Created // 🤖 AI-Generated
🏆Winner @ #MetSoc25 on June 24
Details👉 tinyurl.com/dh37xt78
📩info.emn@metabolomicssociety.org
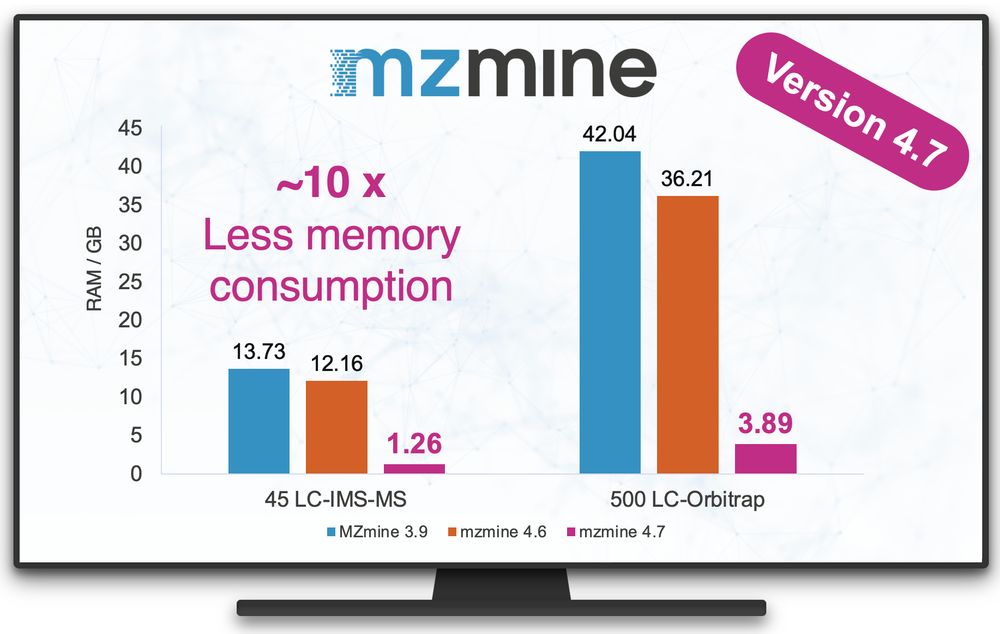

#mzmine 4.7 is now available!
This release brings our most significant improvement in memory efficiency to date, unlocking new capabilities for analyzing large-scale datasets.
Join us for a live software demo at our booth today and tomorrow at 12:00/noon during #ASMS2025

Today is the registration deadline for our bioML symposium on Aug 21-22. Last chance to secure your spot! More info at tinyurl.com/bioMLPrague25

⚛️ @pluskal-lab.org from IOCB Prague, together with his student @roman-bushuiev.bsky.social and colleagues from #CIIRC CTU, Josef Šivic and @anton-bushuiev.bsky.social, have developed a machine learning model called #DreaMS – which accelerates the analysis of previously unknown molecules.
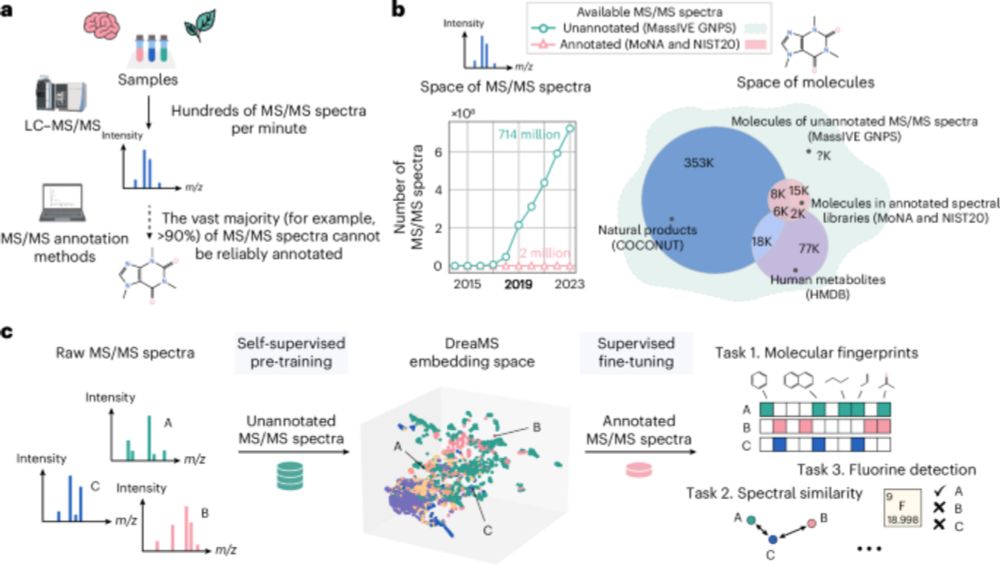
Mass spectrometry is a key method to discover and identify molecules in biological and environmental samples. Yet, >90% of mass spectra remain hard to interpret. In our recent paper, we present DreaMS — a foundation model to interpret mass spectra of small molecules.
www.nature.com/articles/s41...
This paper represents a great effort by @roman-bushuiev.bsky.social and his brother @anton-bushuiev.bsky.social. The DreaMS foundation model for mass spectra of small molecules now opens lots of avenues for possible downstream applications. It might be a game changer for computational metabolomics.

The Mass Spectrometry Query Language (MassQL) is an open-source language for instrument-independent searching across mass spectrometry data for complex patterns of interest via concise and expressive queries without the need for programming skills.
www.nature.com/articles/s41...

Join us for a power-packed software workshop where we’ll be breaking boundaries in multi-vendor MS analysis using #mzmine at #ASMS2015
Led by experts @robinschmid.bsky.social and @ansgarkorf.bsky.social
𝗡𝗼 𝘀𝗶𝗴𝗻–𝘂𝗽 𝗿𝗲𝗾𝘂𝗶𝗿𝗲𝗱, 𝗷𝘂𝘀𝘁 𝗱𝗿𝗼𝗽 𝗯𝘆 𝘁𝗵𝗲 𝗯𝗼𝗼𝘁𝗵 𝗮𝘁 𝟭𝟮 𝗺𝗶𝗱𝗱𝗮𝘆.
See you there! 👋

Get Ready! Only 3 weeks to go until #ASMS2025!
Our team will be showcasing our latest developments in #mzmine at this year’s conference.
Stop by Booth 210 for live demonstrations and insights into how you can streamline your data processing across multiple vendors, adding speed and confidence.

🤝 IOCB Prague is hiring – we are looking for an outstanding Junior Group Leader in Biosciences to join our vibrant scientific community.
See more ► www.uochb.cz/en/open-posi... ⤵️
#openposition #iocbprague #iocb #researchjobs #biosciences
Also thanks to Mélissa Nothias (LMR Naturals by IFF) for her great keynote lecture!
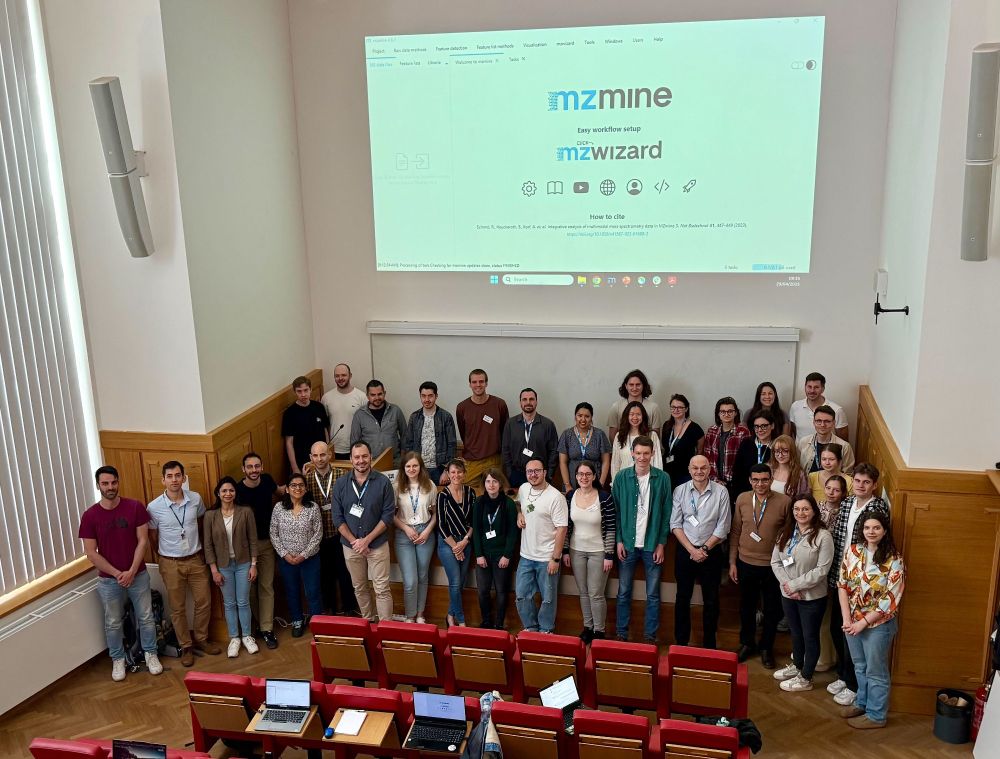

Another year, another @mzmine.bsky.social workshop at
@iocbprague.bsky.social! Both new and advanced users are learning about the latest mzmine features from our amazing instructors @ansgarkorf.bsky.social, Josh Smith,
@titodamiani.bsky.social, and @roman-bushuiev.bsky.social.

Back in July 2023 we organized a small bioML symposium at @iocbprague.bsky.social, and it turned out to be a very pleasant and successful event. This summer we are following up with a great line-up of speakers. Please register for free, deadline May 30. tinyurl.com/bioMLPrague25