


Had an informative and enjoyable time at NeurIPS and MLSB. It was great to see my students present their work and catch up with other CPCB students, both past and present.



Had an informative and enjoyable time at NeurIPS and MLSB. It was great to see my students present their work and catch up with other CPCB students, both past and present.

Excited to be at the first independent @workshopmlsb.bsky.social

If you like to sample from the Boltzmann distribution and are in San Diego for NeurIPS, be sure to check out Rishal's (@rishalchich.bsky.social) poster (#2110). Great work with Nick Boffi (@nmboffi.bsky.social) and Jacky Chen.
neurips.cc/virtual/2025...
arxiv.org/abs/2507.00846
Not true - random number generators aren't biased by molecular weight...
🚨To accommodate the addition of EuroMLSB, we have extended the submission deadline to October 1, 2025 11:59pm AoE.
Find information on paper guidelines at mlsb.io. Submissions will be made through CMT.
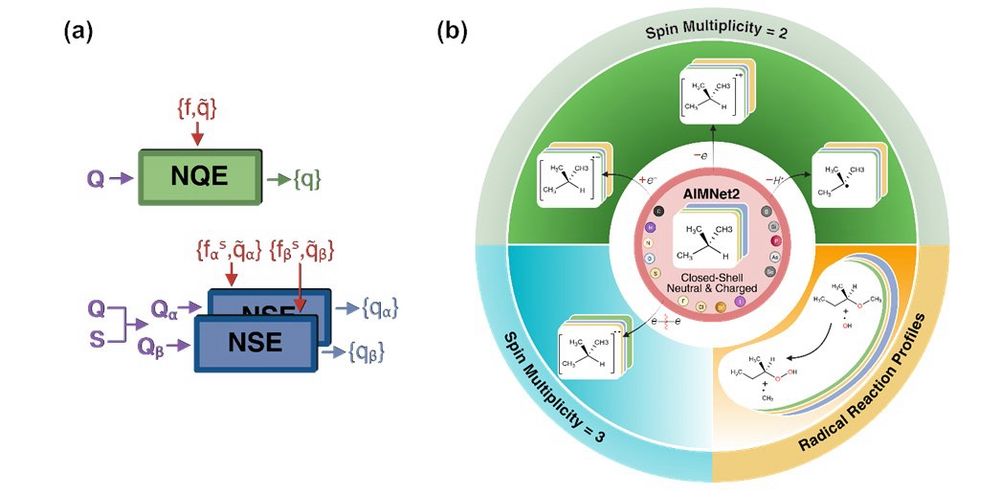
Most of MLIPs dont distinguish between spin states, making them unsuitable for open-shell chemistry. We present AIMNet2-NSE (Neural Spin-charge Equilibration), MLIP that incorporates spin-charge equilibration for systems with arbitrary charge and spin. #compchem #skychem
chemrxiv.org/engage/chemr...
Woohoo!

Protein function often depends on protein dynamics. To design proteins that function like natural ones, how do we predict their dynamics?
@hkws.bsky.social and I are thrilled to share the first big, experimental datasets on protein dynamics and our new model: Dyna-1!
🧵
Nooooooooo.... 😲
NIH funding supporting the HMMER and Infernal software projects has been terminated. NIH states that our work, as well as all other federally funded research at Harvard, is of no benefit to the US.
Our new preprint PharmacoForge: Pharmacophore Generation with Diffusion Models is out now! PharmacoForge quickly generates pharmacophores for a given protein pocket that identify key binding features and find useful compounds in a pharmacophore search. Check it out! 🧪 doi.org/10.26434/che...


Was proud and honored to hood Dr. Drew McNutt at the Pitt School of Medicine Diploma Ceremony. Here we are rocking both the blue and gold and tartan colors representing our joint Pitt-CMU CompBio PhD program. Congratulations to Drew and the other @cmupittcompbio.bsky.social graduates!
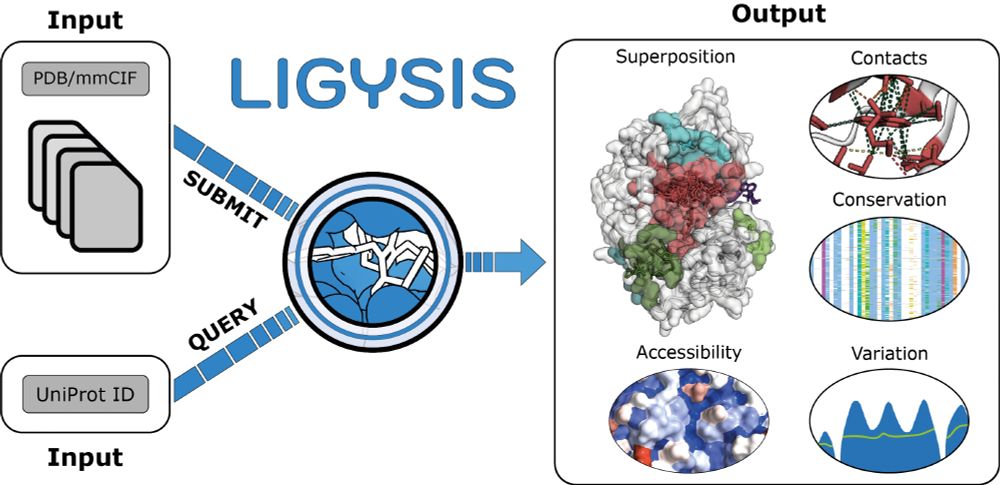
I am #exhilarated to share that our new paper "LIGYSIS-web: a resource for the analysis of protein-ligand binding sites" is now published in @narjournal.bsky.social! After almost two years of development and more than 600 commits, LIGYSIS-web is out!
📜 : tinyurl.com/utges-LIGYSI...

Two students in the Joint Carnegie Mellon-University of Pittsburgh PhD Program in Computational Biology have received honorable mentions from the National Science Foundation’s Graduate Research Fellowship Program. Congratulations to Anamarie Martinez and Emma Flynn!
Read more: tinyurl.com/GRFPPitt
We're recruiting a 3-year postdoc for the Novo Nordisk - Oxford Fellowship programme!
Develop machine learning approaches for fragment library design and experimental optimisation
With @fergusimrie.bsky.social and Charlotte Deane
Job advert: shorturl.at/3l47e
Further details: shorturl.at/u4UkK
Everything is easy in 2D.. - @franknoe.bsky.social

🚀One week left to register for our Symposium on Open Drug Discovery! Join us in Montreal April 7-8 for an exciting program showcasing how open science and AI are driving drug discovery. Some sessions are already sold out, so register now! Register by April 2nd: conscience.ca/symposium2025

Interested in generative modeling and pharmacophores search for SBDD? Check out our talks at #ACSSpring2025
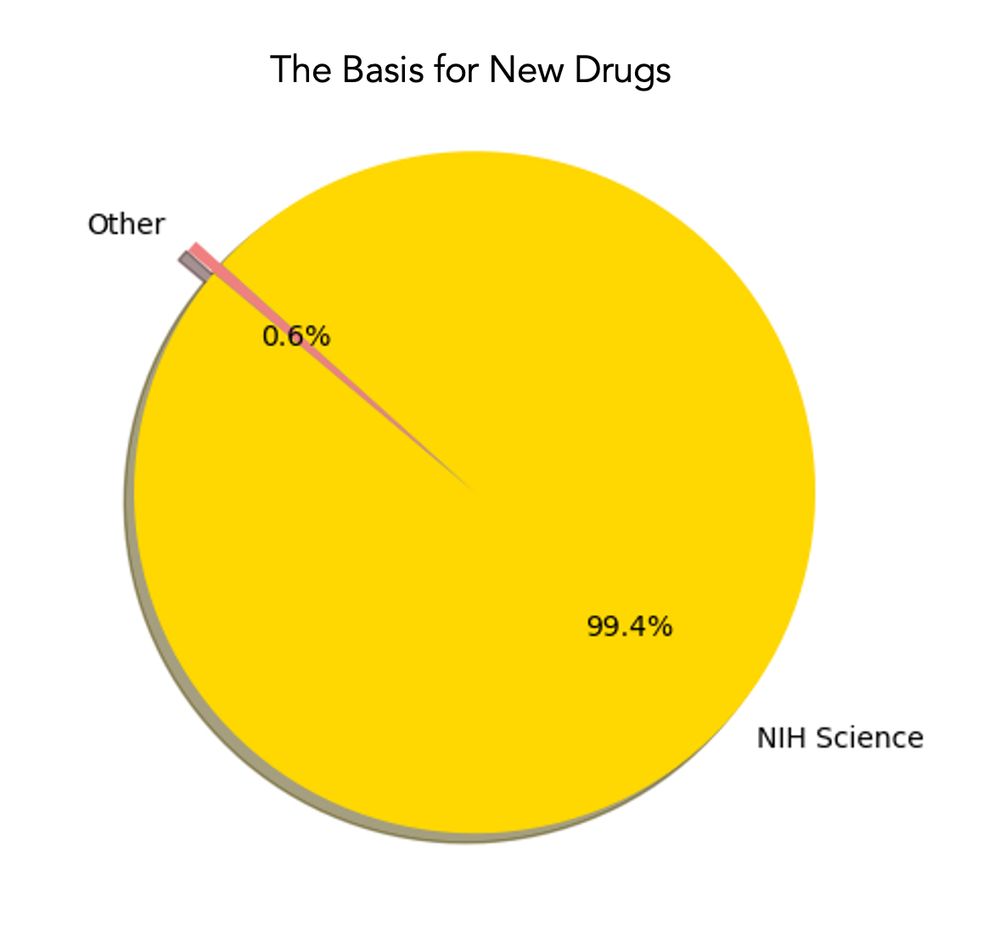
graph of NIH basisfor new drugs
A pie graph worth keeping in mind as the NIH budget plummets jamanetwork.com/journals/jam... for 356 new FDA drugs approved
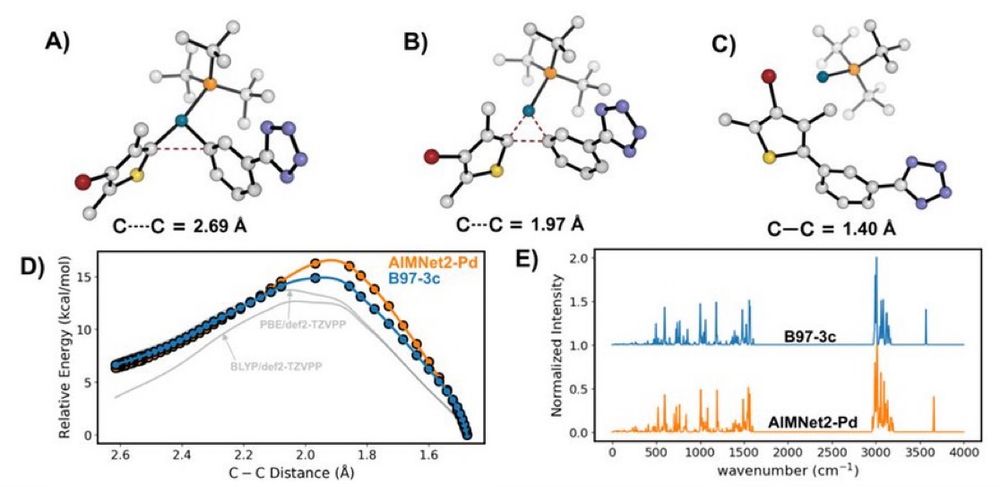
Transferable MachineLearning Interatomic Potential for Pd-Catalyzed Cross-Coupling Reactions
Our latest ChemRxiv preprint, "Transferable #MachineLearning Interatomic Potential for Pd-Catalyzed Cross-Coupling Reactions" Collaboration with @nsf-ccas.bsky.social @gabegomes.bsky.social @bobbypaton.bsky.social chemrxiv.org/engage/chemr... #compchem #chemsky

The CACHE #2 preprint is now online: bit.ly/3DyCmHN Active learning and fragment growing delivered confirmed hits. A citizen scientist using the Fold-it gaming interface designed the top compound! Kudos to Sasha and Madhushika @thesgc.bsky.social at the bench. @conscience-network.bsky.social


@standupforscience.bsky.social
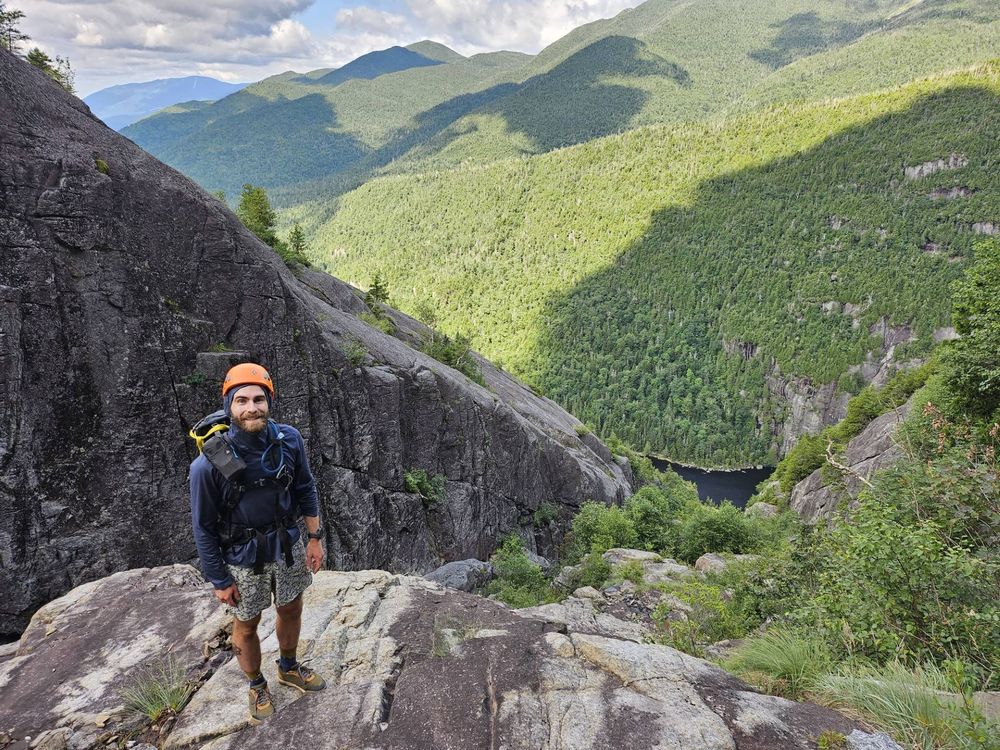

Andrew McNutt on a sola climbing trip in New York

Andrew McNutt poses for a photo with other students in the CPCB program
Andrew McNutt, '24 CPCB graduate, is ready for life's next adventure. Before he begins his career in drug discovery, he's taking a well-deserved break to hike the Appalachian Trail from Georgie to Maine.
Read more: tinyurl.com/AndrewMcNutt

Welcome to the Bluesky account for Stand Up for Science 2025!
Keep an eye on this space for updates, event information, and ways to get involved. We can't wait to see everyone #standupforscience2025 on March 7th, both in DC and locations nationwide!
#scienceforall #sciencenotsilence
I’ll have to read the paper, but that’s a bizarre take given the graph. The issue isn’t the amount of data but the ability to generalize from it - which absolutely requires more innovative algorithms.
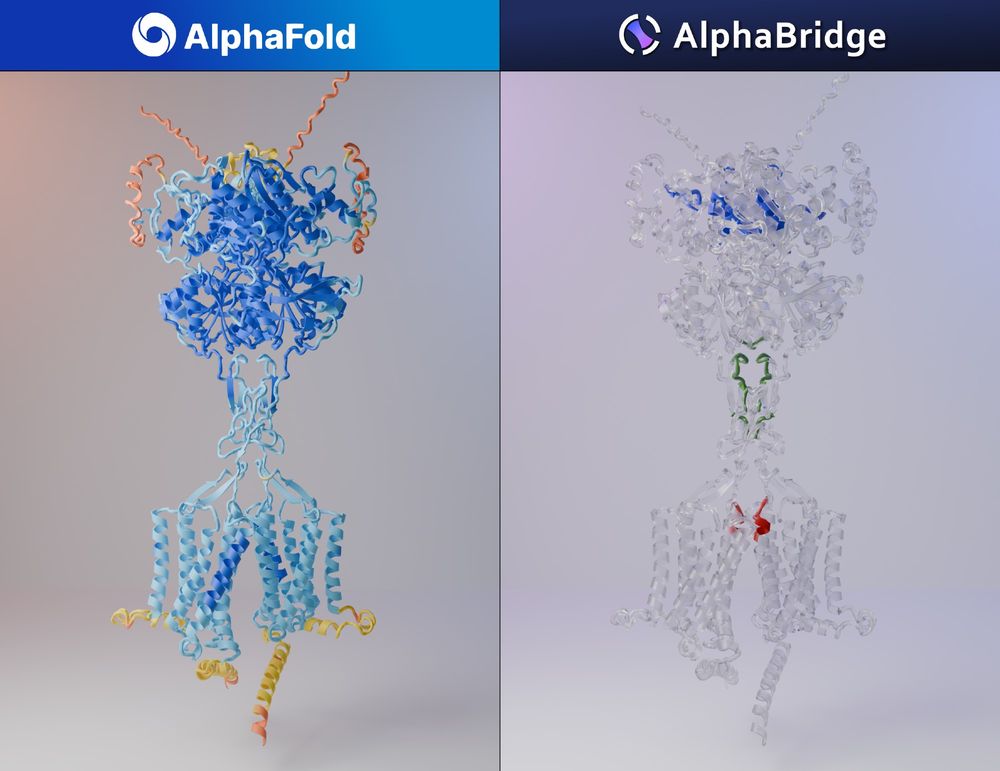

AlphaFold can predict protein, DNA, or RNA complexes.. BUT is it easy to determine the reliable interactions?🤔
➡️ AlphaBridge will identify the most confident interactions, making your analysis easier!
🌐Try it here: alpha-bridge.eu
👀Read more: www.biorxiv.org/content/10.1...
NIH council meeting (NIDCR)scheduled for today, where our R01 grant (3rd percentile) was supposed to be discussed, was postponed indefinitely as a consequence of Trump's executive order which ordered pausing of all communications from all federal agencies. Not good.

Protein structure embeddings reveal undersampled and de novo structure space. (B) First two principal components of mean-pooled ESM3 embeddings colored by helix content determined by DSSP. The indicated dashed guide lines denote visual boundaries of native structure space not sampled (Undersampled) and novel regions of protein structure for this space only observed in samples but not in native structures (De novo). (C) Rasterized visualization of panel B with 16 equally spaced grid squares in each principal component axis. A representative structure from each grid was chosen at random. Empty grid squares indicate the absence of any structure in the enclosed region. De novo alpha helices are shaded along the lower-right diagonal and the structures from CATH which do not have corresponding structures in the samples are shaded along the left and top rims. The structures are displayed in CATH raster plot are given in the Supplementary Information.
Protein backbone diffusion models consistently undersample catalytically important motifs, and oversample idealized helices, raising questions about the appropriateness of these methods in designing starting points for enzyme design & evolution. From www.biorxiv.org/content/10.1...
OPIG is now on Bluesky!
Follow us for updates about the group's latest work, web app updates, and more.
opig.stats.ox.ac.uk