It's time for the 2025 Gibbons Lab Research Roundup!
Yeehaw! 🤠 @isbscience.org
In these dark times, let the joy of scientific discovery and trainee success be your balm ❤️🩹
🧵...
19.12.2025 00:06
👍 18
🔁 5
💬 4
📌 1

2025 ISB Microbiome Symposium - YouTube
The 2025 ISB Microbiome Symposium was a virtual scientific event hosted by the Institute for Systems Biology (ISB) on December 12, 2025. The symposium brough...
If you missed the 2025 @isbscience.org Virtual Microbiome Symposium (theme: how microbial metabolites influence the brain, the immune system, & metabolism), the videos of the talks and the panel discussion are now available here:
youtube.com/playlist?lis...
Thanks to all who helped make it happen!
17.12.2025 16:34
👍 22
🔁 13
💬 1
📌 0

24.09.2025 01:01
👍 1
🔁 0
💬 0
📌 0
Our paper demonstrating that within-species warfare interactions are ecologically important on human skin is now published in Nature Micro! www.nature.com/articles/s41...
30.06.2025 12:26
👍 222
🔁 97
💬 10
📌 3

The paper has the incorrect dynamics for gLV
12.07.2025 17:21
👍 0
🔁 0
💬 0
📌 0
The mbtransfer paper also has data leakage issues. In many of the results they apply DESeq2 Normalization in a pre-processing step (for all samples in a bulk process) and include samples which are later used as forecasting test samples when doing so.
12.07.2025 17:21
👍 1
🔁 0
💬 1
📌 0
Our paper is now open access as well.
29.05.2025 16:23
👍 4
🔁 1
💬 0
📌 0
Call for papers on the clinical microbiome - Nature Microbiology
Nature Microbiology has launched a joint collection on the clinical microbiome with Nature Communications, Nature Medicine and Communications Medicine.
This month's editorial is an invitation to submit papers on clinical and translational aspect of the microbiome.
This collection is a joint effort with @naturemedicine.bsky.social @natcomms.nature.com and @commsbio.nature.com
#MicroSky #MicrobiomeSky 🦠⚕️🧪⚕️
www.nature.com/articles/s41...
06.05.2025 19:37
👍 25
🔁 18
💬 3
📌 1
reviewer 3 or 2?
06.05.2025 22:51
👍 1
🔁 0
💬 0
📌 0

It is a great pleasure to listen to @seppekuehnlab.bsky.social
His talk is about ´Learning microbiome design principles from natural variation ´.
Below, the #liveSketch painting during the seminar and given at the end!
#ArtAndScience #MediationScientifique
05.05.2025 10:30
👍 5
🔁 3
💬 2
📌 0
Openings
Positions available in the Gibson Lab
Several Postdoctoral Fellow openings. All fellows have triple appointment at HMS,BWH, and Broad
- Biological sequence models (theory, design, and application)
- Learning single cell dynamics
- Bacteriotherapy design, “bugs-as-drugs” (using control theory principles)
gibsonlab.io/openings/
17.04.2025 14:02
👍 2
🔁 1
💬 0
📌 0
We are still accepting registrations for this event! Please come join us and hear about cutting edge microbiome research.
17.03.2025 20:05
👍 8
🔁 3
💬 0
📌 0

Planning and describing a microbiome data analysis - Nature Microbiology
We provide guidance on the planning, execution and description of statistical analyses in microbiome studies.
A must-read for any microbiome researcher 🦠📐and a new addition to our #bestpractices series
Planning and describing a microbiome data analysis by @amydwillis.bsky.social and @davidandacat.bsky.social
www.nature.com/articles/s41...
21.02.2025 14:48
👍 58
🔁 26
💬 1
📌 1
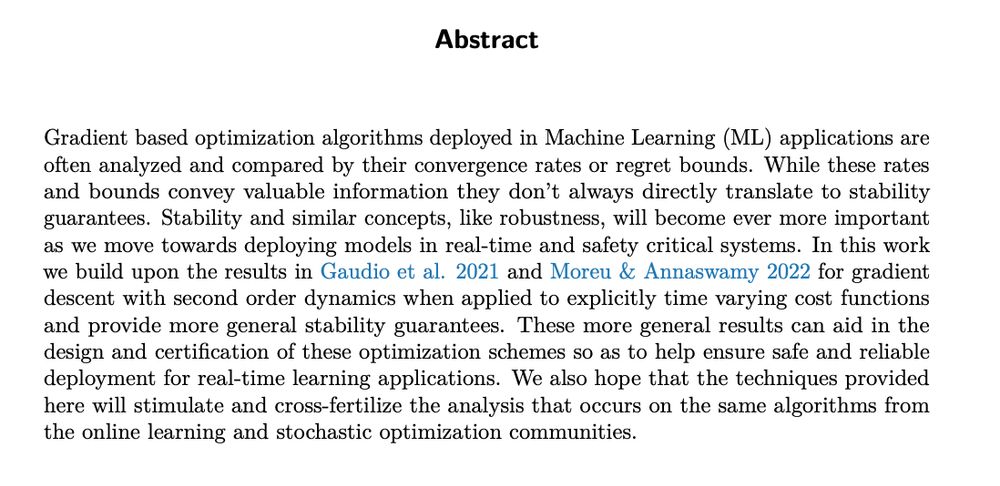

Paper just accepted
Gibson et al. "On the stability of gradient descent with second order dynamics for time-varying cost functions" Transactions Machine Learning Research (2025)
openreview.net/forum?id=Hlz...
07.02.2025 16:35
👍 2
🔁 1
💬 0
📌 0
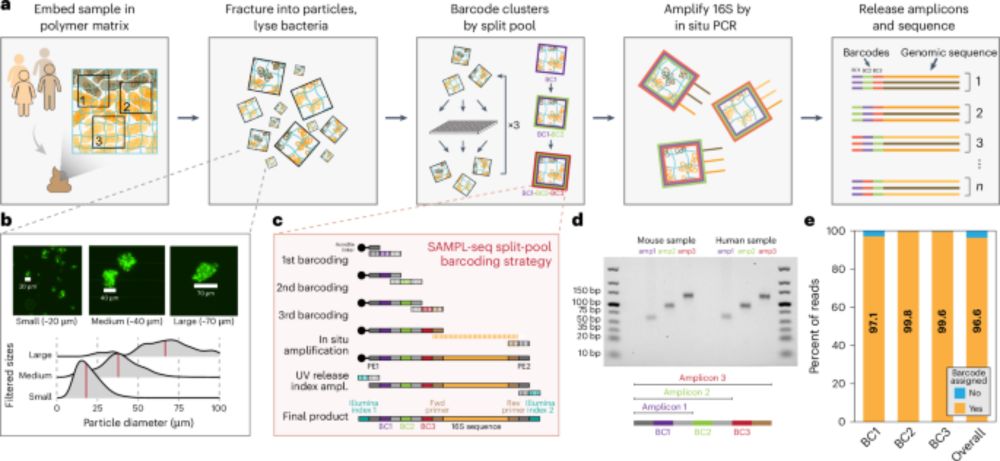
SAMPL-seq reveals micron-scale spatial hubs in the human gut microbiome - Nature Microbiology
Split-And-pool Metagenomic Plot-sampling sequencing (SAMPL-seq) can be applied to complex microbial communities to reveal spatial co-localization of microbes at the micron scale.
Happy to share our new paper in @naturemicrobiol.bsky.social on mapping spatial relationships of the human gut microbiome. We identified distinct spatial hubs between gut bacteria that reflect sub-community assemblies at the micron-scale. Led by Miles Richardson & co.
www.nature.com/articles/s41...
03.02.2025 14:46
👍 89
🔁 36
💬 0
📌 0
Are regret based adaptive control strategies deployable?
arxiv.org/abs/2501.04572
09.01.2025 14:38
👍 0
🔁 0
💬 0
📌 0
It’s easy to think this is about politics, but Ran’s actually just subtweeting my working style
08.12.2024 18:16
👍 13
🔁 2
💬 1
📌 0
As further evidence you should send them the @abrahamgihawi.bsky.social paper that invalidated and lead to the retraction of the Nature paper about a cancer microbiome.
03.12.2024 13:23
👍 5
🔁 0
💬 0
📌 0