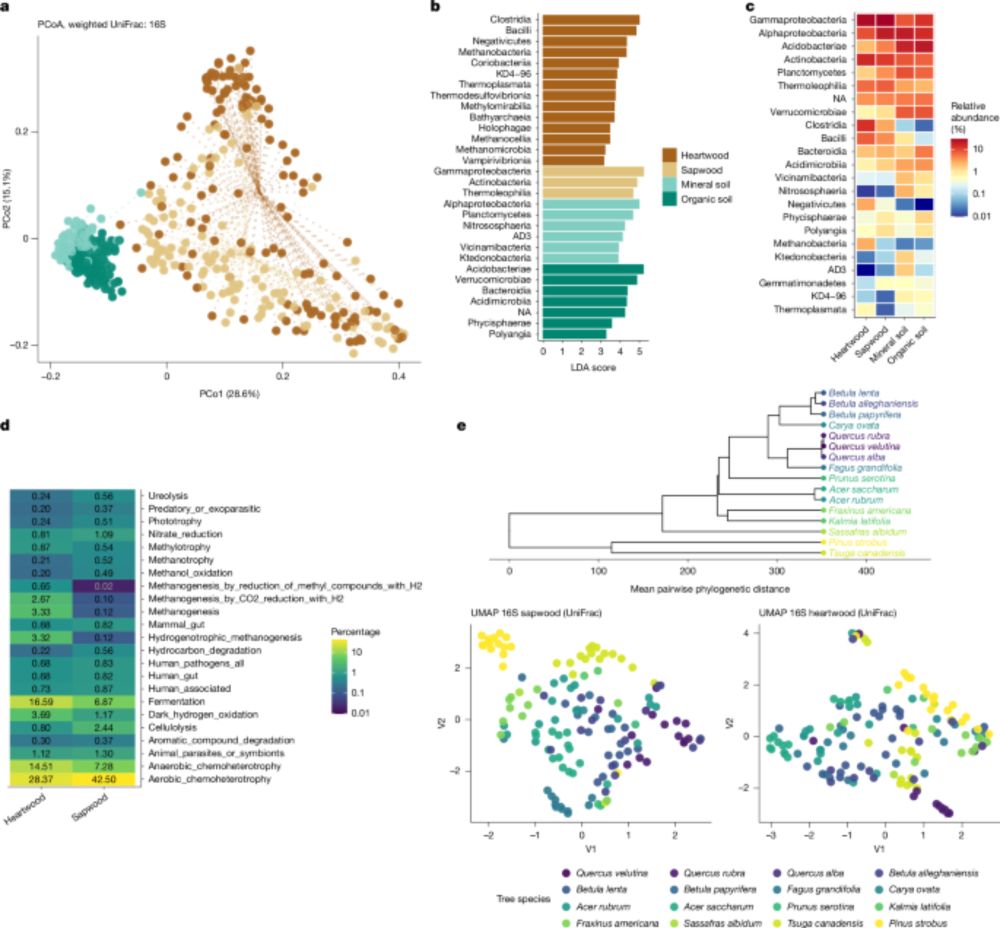
Adult male (small) and female (large) of Armillifer sp. Looks like a big and small crinkle cut french fry, also with paired hooks at the front.
Both photos from this paper:
https://journals.plos.org/plosntds/article?id=10.1371/journal.pntd.0000320

Adult female Linguatula serrata. Looks like a wrinkly tube, a bit wider at the front and tapering to a tail. There are some hooklike bits at the front.
🚨 Don't eat uncooked snake! Your eyes & lungs can be parasitized by pentastomids, which are a CRUSTACEAN. Adults lack so many features that they weren't believed to be arthropods at all until their DNA was sequenced (and sperm morphology, but that was not widely accepted). Yet, they are!
#Crustmas 🧪
21.12.2023 16:38
👍 360
🔁 82
💬 37
📌 36

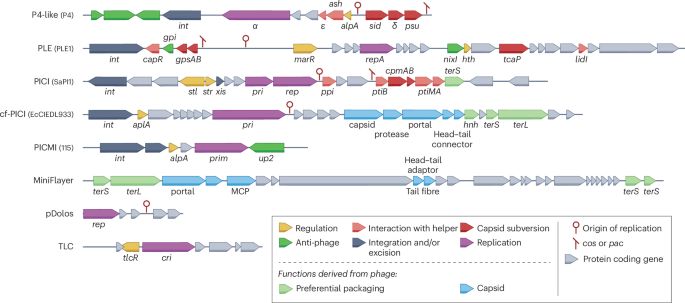
Intracellular competition shapes plasmid population dynamics
From populations of multicellular organisms to selfish genetic elements, conflicts between levels of biological organization are central to evolution. Plasmids are extrachromosomal, self-replicating g...
Hot off the press! Our latest paper led by @fernpizza.bsky.social, understanding how plasmids evolve inside cells. These small, self-replicating DNA circles live inside bacteria and carry antibiotic resistance genes, but also compete with one another to replicate. 1/
www.science.org/doi/10.1126/...
20.11.2025 21:42
👍 437
🔁 199
💬 11
📌 18
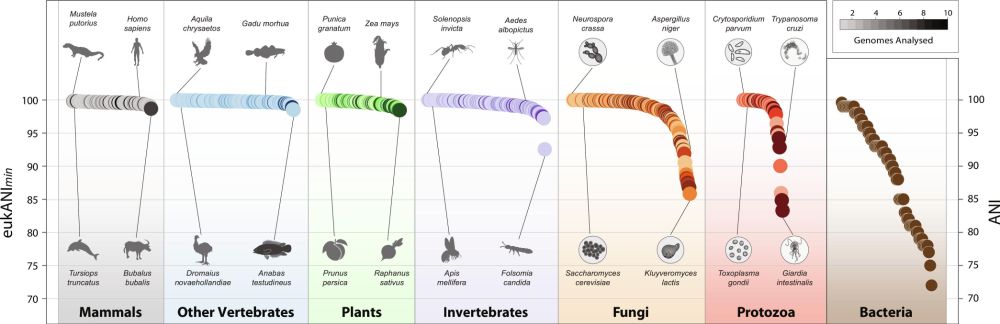
Average nucleotide identity — the backbone of modern ecological genomics - Nature Reviews Genetics
In this Journal Club, Luis Orellana recalls a 2005 publication by Konstantinidis and Tiedje that introduced average nucleotide identity as a sequence-based metric to determine the relatedness between ...
The average nucleotide identity (ANI) underpins how we map microbial diversity, compare species, and connect genomes to ecology.
I wrote a short piece reflecting on the discovery and significance of this metric (and really enjoyed digging into the context and story behind it!) #microsky 🧬
30.10.2025 21:11
👍 58
🔁 22
💬 1
📌 2
C. elegans is a real animal and we set out to understand how it comes to have its distinctive biogeography. Its ancestral center of diversity is in the higher elevation forests of Hawaii. Its closest relatives are spread across east Asia. Did they travel from Asia? [Preprint 🧵]
24.09.2025 20:33
👍 168
🔁 79
💬 5
📌 8
Heads up: ignore samtools dot org, similarly minimap2 dot com and likely others. It's owned by a known phishing site and while the binaries they offer look valid currently (but note they may be serving us different binaries to others), that could change.
Ie: it's not us (Samtools team)! Be warned
15.09.2025 08:40
👍 146
🔁 127
💬 2
📌 5
tgv 0.1.0 release: github.com/zeqianli/tgv
- Rich CIGAR and base visualization
- Allele frequency visualization
- VCF and BED file support
- Mouse dragging and hovering
- Filter alignment
Now 90% of what I need from IGV can be done in the terminal.
Some interesting behind-the-scenes:
07.09.2025 23:47
👍 11
🔁 7
💬 1
📌 0

Oatk: a de novo assembly tool for complex plant organelle genomes. #DeNovoAssembly #OrganelleGenomes #Bioinformatics #GenomeBiology
genomebiology.biomedcentral.com/articles/10....
11.08.2025 10:30
👍 5
🔁 4
💬 0
📌 0

StrainR2 accurately deconvolutes strain-level abundances in synthetic microbial communities. #Metagenomics #StrainLevelAbundance #Bioinformatics
academic.oup.com/bioinformati...
09.08.2025 18:05
👍 2
🔁 2
💬 1
📌 0

Protein language models reveal evolutionary constraints on synonymous codon choice
Evolution has shaped the genetic code, with subtle pressures leading to preferences for some synonymous codons over others. Codons are translated at different speeds by the ribosome, imposing constrai...
Protein language models reveal evolutionary constraints on synonymous codon choice
#rnasky #microsky "cotranslational localization and translational accuracy, more than cotranslational protein folding, are major drivers of selective pressure on codon choice" in yeast here 💫
doi.org/10.1101/2025...
09.08.2025 18:31
👍 3
🔁 1
💬 0
📌 0
Announcing taxburst, an update of the Krona software for taxonomy exploration
Announcing taxburst for metagenome taxonomy!
taxburst v0.3.0 is now released - this is an update of the Krona visualization system for microbiome/metagenome taxonomy analyses. Enjoy!
08.08.2025 14:19
👍 26
🔁 17
💬 0
📌 0

Scaling down protein language modeling with MSA Pairformer
Recent efforts in protein language modeling have focused on scaling single-sequence models and their training data, requiring vast compute resources that limit accessibility. Although models that use ...
Excited to share work with
Zhidian Zhang, @milot.bsky.social, @martinsteinegger.bsky.social, and @sokrypton.org
biorxiv.org/content/10.1...
TLDR: We introduce MSA Pairformer, a 111M parameter protein language model that challenges the scaling paradigm in self-supervised protein language modeling🧵
05.08.2025 06:29
👍 97
🔁 43
💬 1
📌 1
GitHub - lh3/longdust: Identify long STRs, VNTRs, satellite DNA and other low-complexity regions in a genome
Identify long STRs, VNTRs, satellite DNA and other low-complexity regions in a genome - lh3/longdust
Fun new tool from Heng Li. Thinking maybe I can use this to help find plasmid replication gene correlated repeat regions - though he specifically mentions it's not for tandem repeat regions. Hmm. 🖥️🧬
github.com/lh3/longdust
01.08.2025 11:34
👍 7
🔁 2
💬 0
📌 0
A bit late to joining the Bluesky party, but it's great to see all the amazing scientists who are on this platform! Looking forward to connecting with all of you here (on twitter as @niranjantw ... so keeping the handle consistent).
28.04.2025 01:13
👍 5
🔁 2
💬 2
📌 0
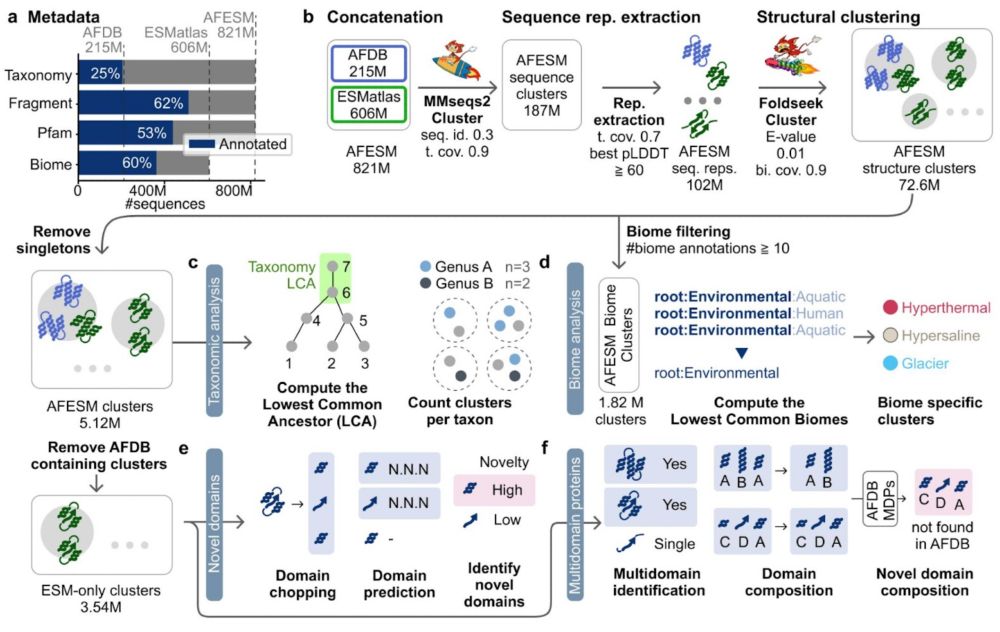
AFESM: a metagenomic guide through the protein structure universe! We clustered 821M structures (AFDB&ESMatlas) into 5.12M groups; revealing biome-specific groups, only 1 new fold even after AlphaFold2 re-prediction & many novel domain combos. 🧵
🌐 afesm.foldseek.com
📄 www.biorxiv.org/content/10.1...
27.04.2025 00:13
👍 141
🔁 71
💬 4
📌 4

A pack of stickers with the logo of our new tool, K-mer Fast Counter a.k.a. KFC
I'll be presenting our work on hyper-k-mers at #RECOMB today at 10:40 KST!
You can get a sneak peek at the slides here: igor.martayan.org/slides-recom...
Come say hi if you'd like to chat, or just get one of these cute stickers!
26.04.2025 22:23
👍 18
🔁 4
💬 1
📌 0
Assemblies of long-read metagenomes suffer from diverse errors https://www.biorxiv.org/content/10.1101/2025.04.22.649783v1
25.04.2025 00:46
👍 23
🔁 11
💬 0
📌 1
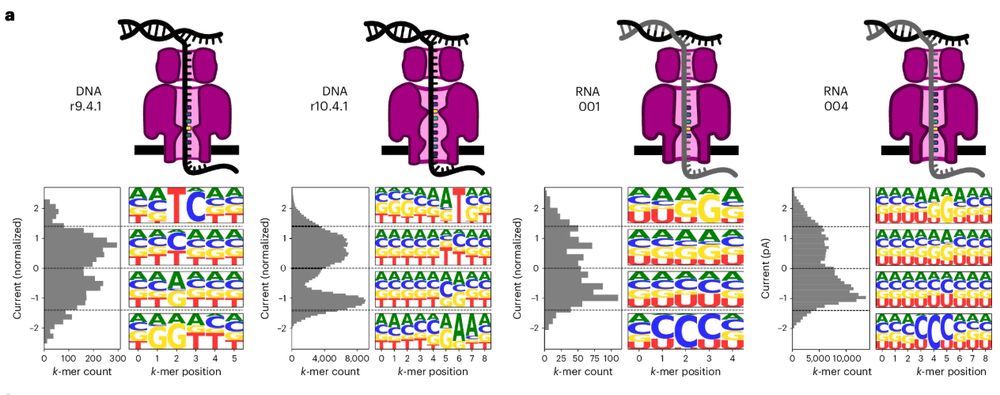
Uncalled4: a toolkit for nanopore signal alignment, analysis and visualization of DNA and RNA modifications.
www.nature.com/articles/s41...
28.03.2025 17:23
👍 47
🔁 26
💬 1
📌 1
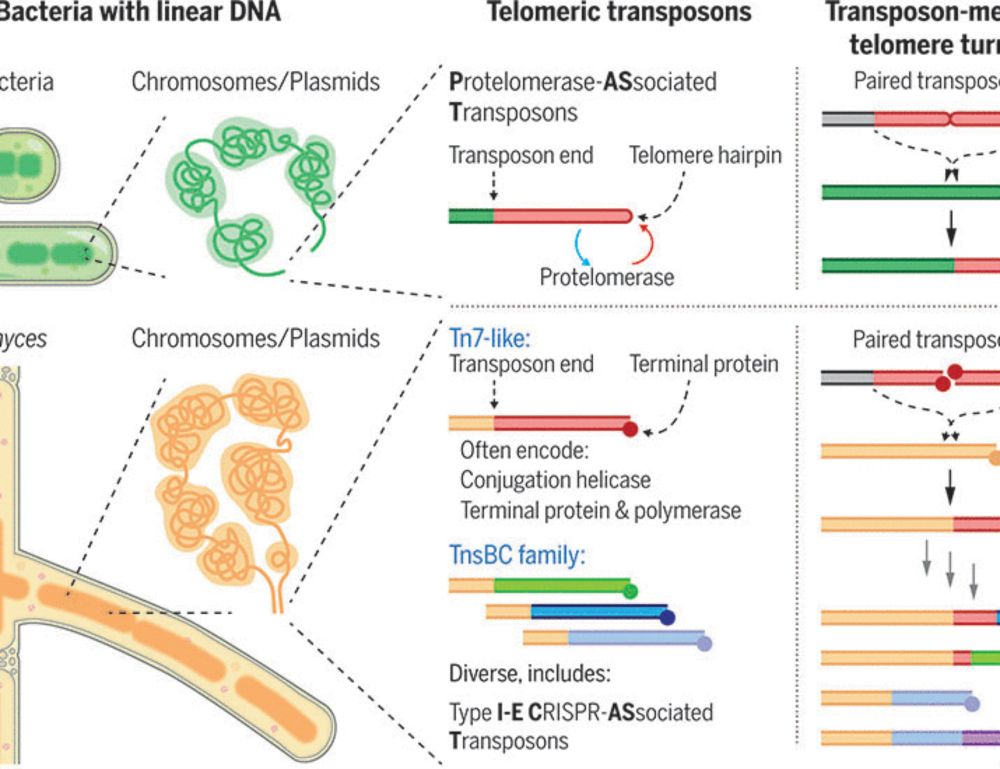
Telomeric transposons are pervasive in linear bacterial genomes
Eukaryotes have linear DNA, and their telomeres are hotspots for transposons, which in some cases took over telomere maintenance. We identified several families of independently evolved telomeric tran...
wow, telomeric transposons in bacteria with linear chromosomes! (of course this was first figured out in flies, inc by Bob Levis, who i was happy to see few days ago at the fly meeting). 🪰
www.science.org/doi/10.1126/...
www.sciencedirect.com/science/arti...
www.sciencedirect.com/science/arti...
27.03.2025 20:55
👍 62
🔁 36
💬 0
📌 1
GitHub - rrwick/condaenvlist: a simple tool for listing conda environments with descriptions
a simple tool for listing conda environments with descriptions - rrwick/condaenvlist
Do you (like me) create a bunch of conda environments, then later forget what they're for, when they were last updated, or which tools are in them?
If so, you might this little project: github.com/rrwick/conda...
27.03.2025 04:34
👍 78
🔁 40
💬 1
📌 1

GitHub - yangao07/longcallD: A local-haplotagging-based small and structural variant caller
A local-haplotagging-based small and structural variant caller - yangao07/longcallD
longcallD is a new variant caller for genomic long reads. It jointly calls phased small and structural variants. Single binary, one command line for the whole process. Comparable accuracy to mainstream callers. Great work by Yan Gao. github.com/yangao07/lon...
24.03.2025 16:53
👍 105
🔁 49
💬 3
📌 3

Interested in bacterial genomes?
Hundreds of thousands, even millions?
All annotated, taxonomically classified, integrated with metadata.
Easily searchable, viewable, downloadable, in sync with #AllTheBacteria.
Then BakRep is for you! Poster P-CM-102 @vaam-microbes.bsky.social #VAAM25
25.03.2025 09:45
👍 18
🔁 9
💬 2
📌 0