Immensely grateful to the Schmidt Science Fellowship, my mentors, and all the wonderful & inspiring people I have worked with over the past years.
Immensely grateful to the Schmidt Science Fellowship, my mentors, and all the wonderful & inspiring people I have worked with over the past years.
Excited to be pivoting from molecules to cells for my Schmidt Science Fellowship, advised by Emma Lundberg and Wah Chiu!
I’m looking forward to building ML models that better reflect how molecules & cells look in real life 🔬
Huge thanks to my co-authors @cschneider.bsky.social, Lewis Chinery, Charlotte Deane @opig.stats.ox.ac.uk!
More information about the paper: bsky.app/profile/alis...
Our paper on generalizable antibody-antigen binding affinity prediction has been featured on the cover of the August Issue of @natcomputsci.nature.com! 📔🎉

Yellow and orange antibody binding to purple and blue membrane proteins
🚨Our August issue is now live and includes research on antibody-antigen binding, molecular screening for zeolite synthesis, psychological experiments with LLMs, and much more!
www.nature.com/natcomputsci...
Out now! @alissahummer.com @opig.stats.ox.ac.uk
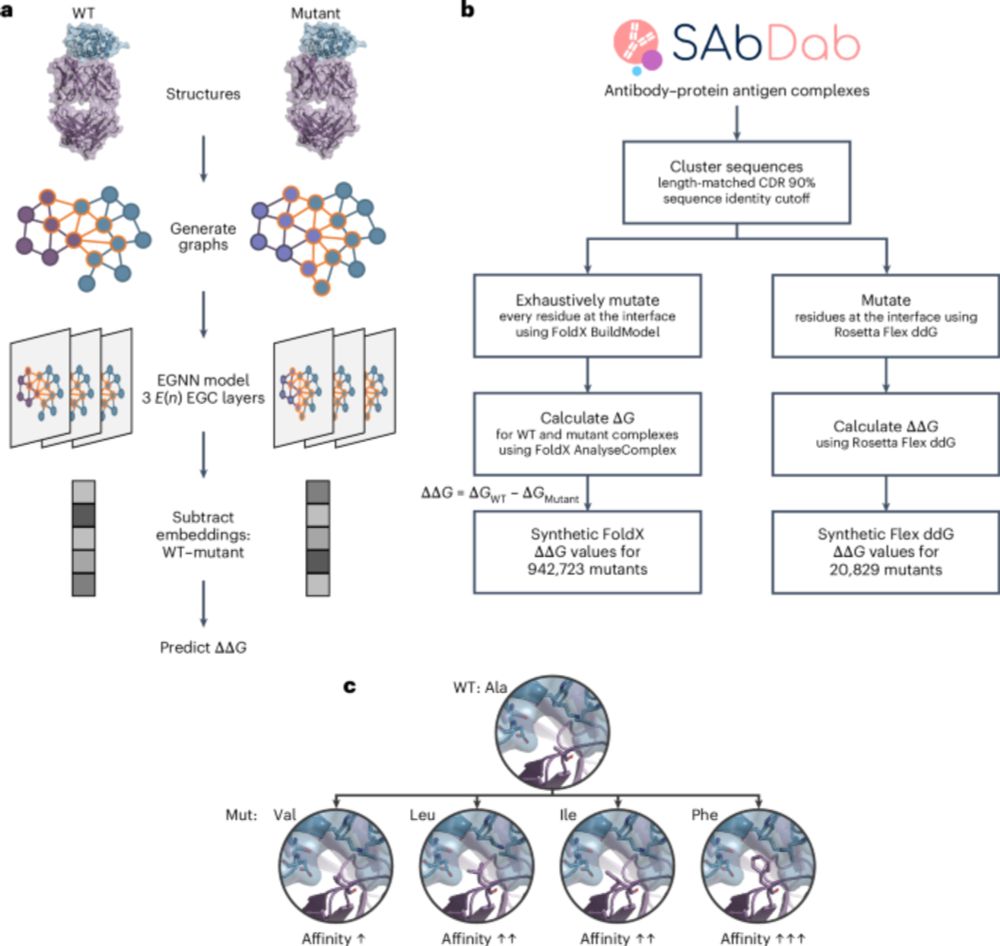
and colleagues present Graphinity, a method to predict change in antibody-antigen binding affinity (∆∆G). Also featuring synthetic datasets of ~1 million FoldX-generated and >20,000 Rosetta Flex ddG-generated ∆∆G values!
www.nature.com/articles/s43...
And thank you to my co-authors @cschneider.bsky.social, Lewis Chinery, and Charlotte Deane – @opig.stats.ox.ac.uk!
Thank you very much to the reviewers and editors who made this work stronger!

The Graphinity code and synthetic datasets are publicly available at the following links.
GitHub: github.com/oxpig/Graphi...
Zenodo (code): doi.org/10.5281/zeno...
Zenodo (data): doi.org/10.5281/zeno...
OPIG (data): opig.stats.ox.ac.uk/data/downloa...
Since the release of our preprint, we have made key updates including:
📊 Generation of a second synthetic dataset using Rosetta Flex ddG (20,829 ΔΔG values)
🕸️ Evaluation of additional ML architectures (incl. FLAML, CNN, Rotamer Density Estimate, Equiformer)
Antibody-antigen binding affinity lies at the heart of therapeutic antibody development.
We show that orders of magnitude more data will be needed to unlock generalizable ΔΔG prediction. Our findings provide a lower bound on data requirements to inform future method development & data collection.
Our work exploring the ability of and requirements for ML to predict the effects of mutations on antibody-antigen binding affinity (ΔΔG) is out now in @natcomputsci.nature.com!

What is the status of nucleic acid structure prediction? Our analysis of CASP16 (doi.org/10.1101/2025...) reveals human expertise is still necessary for the most accurate prediction, but accuracy still heavily relies on templates; having seen a similar structure already.

See the full analysis, led by @rachael-kretsch.bsky.social, in our preprint: www.biorxiv.org/content/10.1...

It was great to be involved in the evaluation of nucleic acid structure prediction in CASP16! 🧬
RNA modeling remains challenging for deep learning, esp. in the absence of templates and for long-range tertiary/quaternary interactions. Encouraging signs from deep evolutionary data though.
Thank you so much, Nick!
If you’re interested in building the data & ML to create molecular-resolution Virtual Cell Models, I would love to chat 🧫💻
And thank you to the @schmidtsciences.bsky.social program for encouraging and supporting disciplinary pivots to enable greater impact for our science!
I'm so excited to join this interdisciplinary community as a 2025 Schmidt Science Fellow!
After years behind a keyboard, I will be pivoting toward the wet lab. To unlock the true potential of ML for biology/biomedicine, we need high-quality data and robust evaluation 🔬🧫🧪
A huge thank you to all my mentors & lab-mates who have shaped my scientific journey and supported my pivots so far!
Special thanks to Charlotte Deane & @opig.stats.ox.ac.uk, @deboramarks.bsky.social, Madan Babu, @pstansfeld.bsky.social, @rdaslab.bsky.social

AntiFold, our antibody inverse folding model, has been published at Bioinformatics Advances. Work led by @magnushoie.bsky.social & @alissahummer.com.
Paper: academic.oup.com/bioinformati...
Webserver: opig.stats.ox.ac.uk/webapps/anti...
Codebase available on Github: github.com/oxpig/AntiFold