
We’re recruiting a postdoc to start at the end of this year or early next year! We use cryo-EM to uncover the structural basis of key chromatin events using native complexes isolated from human cells or assembled in Xenopus (frog) egg extracts!

We’re recruiting a postdoc to start at the end of this year or early next year! We use cryo-EM to uncover the structural basis of key chromatin events using native complexes isolated from human cells or assembled in Xenopus (frog) egg extracts!

My awesome colleague, @yarimura.bsky.social, is recruiting a postdoc to start at the end of 2026 or early 2027! His lab uses cryo-EM to uncover structural basis of key chromatin events using native complexes isolated from human cells or Xenopus egg extracts!
careers-fhcrc.icims.com/jobs/30706/p...
Thank you for the repost, Harmit😆
Thank you for your kind words, CJ! It was a big relief to me!
Thank you, Peter!




@jhigginswriter.bsky.social wrote an article about my lab and our frog 🐸 at the Hutch! Thank you!

Dr. @yarimura.bsky.social brings frogs back to Fred Hutch after a long absence to study the structure of DNA-linked molecular complexes that change during the cell cycle and malfunction in cancer and other diseases. bit.ly/3J8S5Ae

Looking for your next step in fly genetics? The Rajan Lab at Fred Hutch is hiring a staff scientist + postdoc for projects on fat–brain signals, immunity, and brain aging. Stable, long-term role in a collaborative lab. Apply: careers-fhcrc.icims.com/jobs/30062/j...

My awesome Drosophila colleague Akhila Rajan at Fred Hutch (Seattle) is recruiting both a staff scientist and a postdoc to study fat–brain communication, innate immunity, mitochondrial signaling, and brain senescence. Great team, great environment. Apply here: careers-fhcrc.icims.com/jobs/30062/j...

New online: Computational design of sequence-specific DNA-binding proteins
One more movie!
Using the RL algorithm (from pnas.org/doi/10.1073/...),
@masaashimazoe.bsky.social analyzed our nucleosome tracking data and classified them by their mobility.
Hotter color = faster motion (Slow🔵→ 🟢 → 🟡 → 🟠 Fast)

In new study led by @bdadonaite.bsky.social, we measure how spike mutations affect function & antigenicity of spike of KP.3.1.1 strain of SARS-CoV-2.
Sheds light on how key neutralizing epitopes are changing & importance of RBD up/down motion.
www.biorxiv.org/content/10.1...

WORK!
I currently have 5 (!) open positions for postdocs/PhD students in our CDlab for projects:
- Nanopore protein sequencing
- Archaeal CDV cell division
- Microfluidics for synthetic cells
- Nuclear Pore Complex
- Origami mimics of peroxisomes
Please apply! ceesdekkerlab.nl/come-join-us/
RT=👍
The peer-reviewed VOR for the MagIC-cryo-EM paper by @yarimura.bsky.social and @1001hak.bsky.social is finally published!
Congrats Yasu!

🧬🎉Thrilled to share our new chromatin remodeling study! We reveal three states of human CHD1 and identify a novel "anchor element" that interacts with the acidic patch—conserved among remodelers. Our structures clarify mechanisms of remodeler recruitment! Link: authors.elsevier.com/a/1l2ik3vVUP...
Excited to share our preprint on the molecular architecture of heterochromatin in human cells 🧬🔬w/ @jpkreysing.bsky.social, @johannesbetz.bsky.social,
@marinalusic.bsky.social, Turoňová lab, @hummerlab.bsky.social @becklab.bsky.social @mpibp.bsky.social
🔗 Preprint here tinyurl.com/3a74uanv
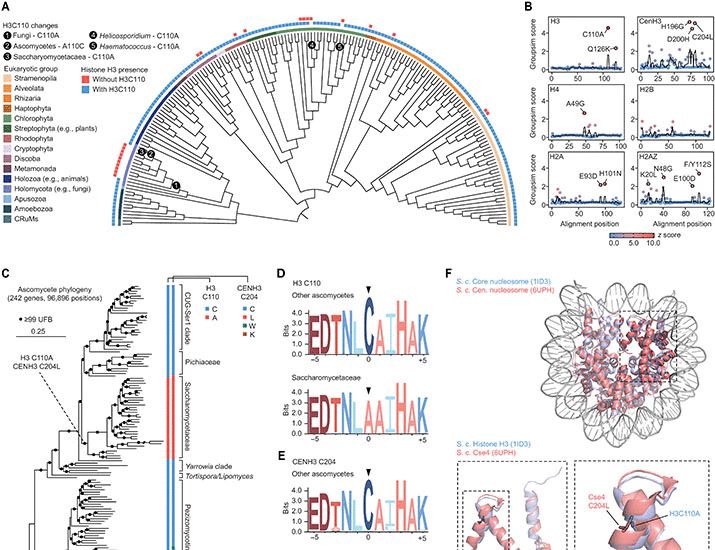
Ever wondered what the lonely histone H3C110 does? We found that it’s lost in many fungi. Adding it back boosts histone H3 copper reductase activity, rescues iron defects and modulates lifespan, suggesting an evolutionary tradeoff between these effects.
www.science.org/doi/10.1126/...
New preprint on 3D heterochromatin architecture in human cells! Great collab with @sergiocruzleon.bsky.social & @johannesbetz.bsky.social from @hummerlab.bsky.social, @marinalusic.bsky.social & the Turoňová lab. Many thanks to my supervisor @becklab.bsky.social. bioRxiv: tinyurl.com/3a74uanv 🧵👇
🔬 New from the Farnung Lab: We established a fully in vitro reconstituted chromatin replication system and report the first cryo-EM snapshots of the human replisome engaging nucleosomes. Brilliant work by @felixsteinruecke.bsky.social with support from @jonmarkert.bsky.social! tinyurl.com/replisome
Our new paper is out@ScienceAdvances👇
www.science.org/doi/10.1126/...
🧬Our Repli-Histo labeling marks nucleosomes in euchromatin and heterochromatin in live human cells.
🔍 @katsuminami.bsky.social et al. have developed a chromatin behavior atlas within the nucleus. 1/2
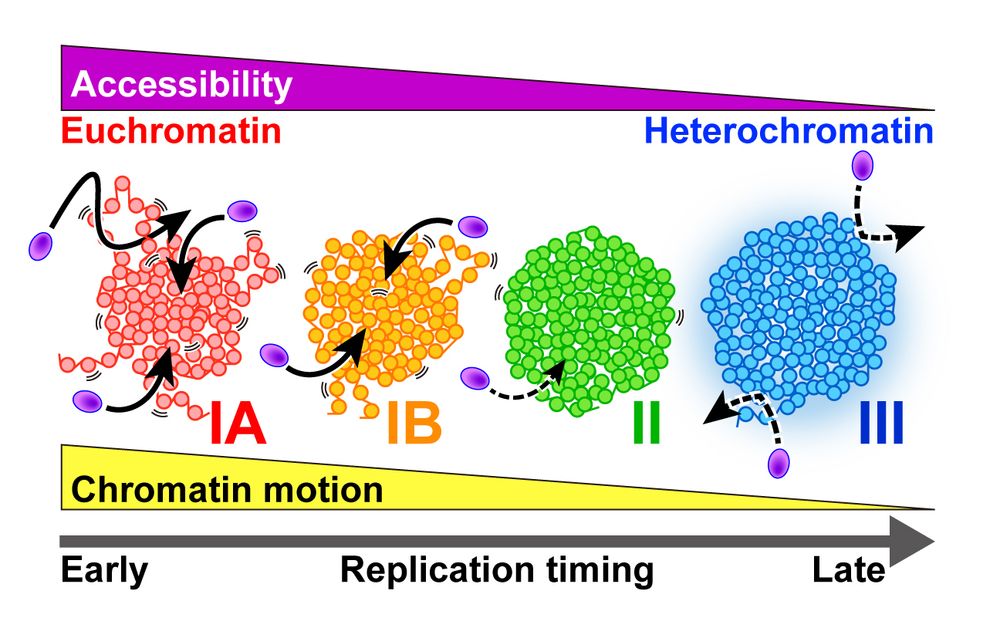
We quantitatively measured nucleosome motion across the 4 chromatin classes, from euchromatin to heterochromatin.
🧪 More euchromatic regions (i.e., earlier-replicated regions) show greater nucleosome motion.
🎉 Congrats @katsuminami.bsky.social @kakonakazato.bsky.social and all collaborators!
2/2

Proud of this work, led by the amazing Jeannette Tenthorey (now Assistant Professor at UCSF), demonstrating the adaptive power of single indel mutations in host-virus arms races, but a little sad that this is the coda to her amazing postdoc in the Malik & Emerman labs.
www.cell.com/cell-genomic...

First post on bsky. We've updated our preprint about the change in the histone H3 variant on chromatin during Zygotic Genome Activation. Thanks to new experiments by Anusha, it seems like our initial, simplistic idea that the embryo "ran out" of H3 was not right.
www.biorxiv.org/content/10.1...
In a new Science study, cryo–electron tomography captures the in-cell architecture of the mitochondrial respiratory chain, illuminating how the coordinated action of molecular machines drives life’s fundamental energy conversion.
Learn more in this week's issue: scim.ag/3FA3Ygq
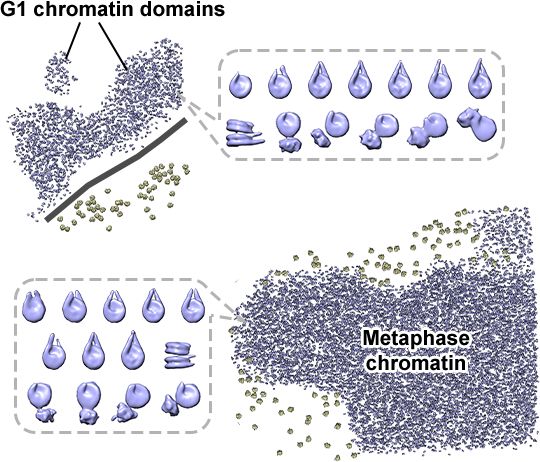
Cryo-ET visualization of chromatin at the nucleosome level in RPE-1 cells arrested in G1 phase and metaphase.
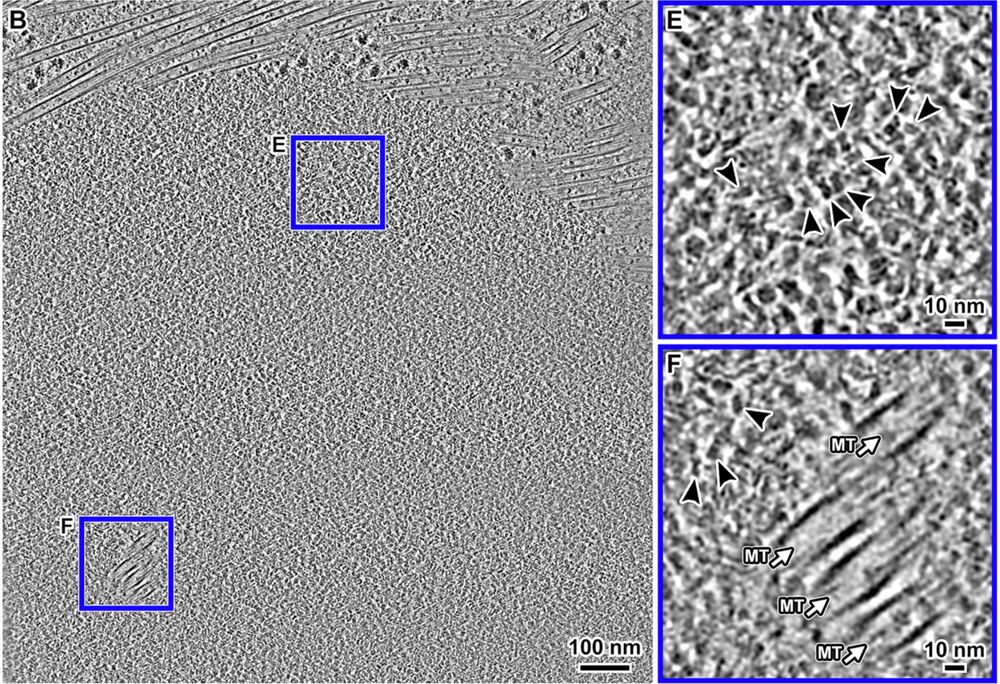

Cryotomograms of a metaphase RPE-1 cell.
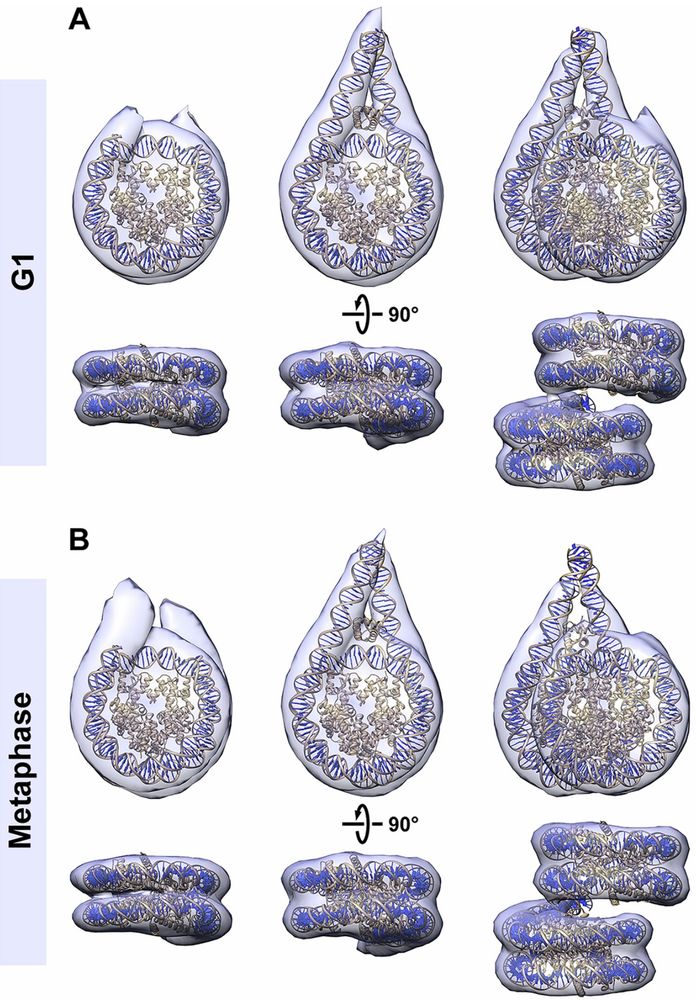
3-D class averages of mononucleosomes and ordered stacked dinucleosomes in G1 and metaphase.
Happy that Jon Chen @jonchenjk.bsky.social's cryo-ET and confocal study of RPE-1 chromatin in situ is online www.embopress.org/doi/full/10.1038/s44318-025-00407-2
Congrats also to coauthors Tingsheng Liu, @shujuncai.bsky.social, Cai Tong Ng, Jian Shi and collaborators Weimei Ruan Uttam Surana.

Wonderful visit to @hutchbasicsci.bsky.social today and thanks to @yarimura.bsky.social for hosting and dinner with the always entertaining @suebiggins.bsky.social and @harmitmalik.bsky.social. A day discussing science with amazing people is unbeatable.
Thank you for visiting us and giving the exciting seminar!
My first, first author paper is now on bioRxiv! 🤩
Linker histone H1 is a liquid-like "glue" condensing chromatin, which revises textbooks! 📖✨
biorxiv.org/content/10.1...
Huge thanks to my PI, @kazu-maeshima.bsky.social, for supervision.
Amazing collab with @rcollepardo.bsky.social’s group!" (1/n)
I am super excited to share that my work at @yifan_ucsf has just been published in Nature🎉 From developing pYC KI system, it was a long journey to understand endogenous protein dynamics. I appreciate the support from FASN team! 🕺Let's dive into the endogenous & dynamic protein world🏊🏄🚣